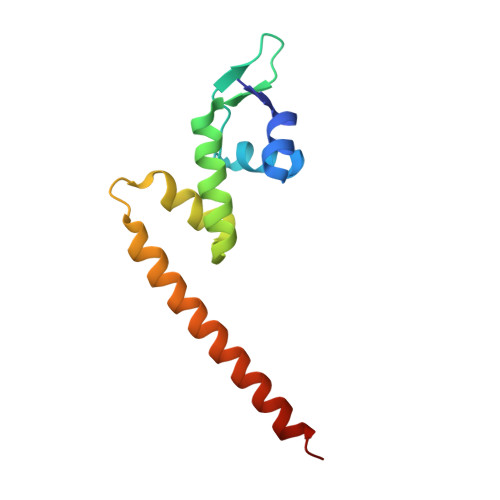
Crystal structure of MtaN, a global multidrug transporter gene activator.
Godsey, M.H., Baranova, N.N., Neyfakh, A.A., Brennan, R.G.(2001) J Biological Chem 276: 47178-47184
- PubMed: 11581256 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M105819200
- Primary Citation Related Structures:
1JBG - PubMed Abstract:
MtaN (Multidrug Transporter Activation, N terminus) is a constitutive, transcriptionally active 109-residue truncation mutant, which contains only the N-terminal DNA-binding and dimerization domains of MerR family member Mta. The 2.75 A resolution crystal structure of apo-MtaN reveals a winged helix-turn-helix protein with a protruding 8-turn helix (alpha5) that is involved in dimerization by the formation of an antiparallel coiled-coil. The hydrophobic core and helices alpha1 through alpha4 are structurally homologous to MerR family member BmrR bound to DNA, whereas one wing (Wing 1) is shifted. Differences between the orientation of alpha5 with respect to the core and the revolution of the antiparallel coiled-coil lead to significantly altered conformations of MtaN and BmrR dimers. These shifts result in a conformation of MtaN that appears to be incompatible with the transcription activation mechanism of BmrR and suggest that additional DNA-induced structural changes are necessary.
- Department of Biochemistry and Molecular Biology, Oregon Health & Science University, Portland, OR 97201-3098, USA.
Organizational Affiliation: