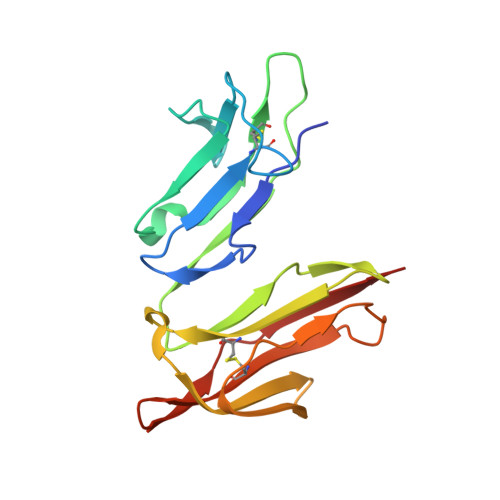
The analysis of the human high affinity IgE receptor Fc epsilon Ri alpha from multiple crystal forms.
Garman, S.C., Sechi, S., Kinet, J.P., Jardetzky, T.S.(2001) J Mol Biology 311: 1049-1062
- PubMed: 11531339 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4929
- Primary Citation Related Structures:
1J86, 1J87, 1J88, 1J89 - PubMed Abstract:
We have solved the structure of the human high affinity IgE receptor, Fc epsilon RI alpha, in six different crystal forms, showing the structure in 15 different chemical environments. This database of structures shows no change in the overall shape of the molecule, as the angle between domains 1 and 2 (D1 and D2) varies little across the ensemble. However, the receptor has local conformational variability in the C' strand of D2 and in the BC loop of D1. In every crystal form, a residue inserts between tryptophan residues 87 and 110, mimicking the position of a proline from the IgE ligand. The different crystal forms reveal a distribution of carbohydrates lining the front and back surfaces of the structure. An analysis of crystal contacts in the different forms indicates regions where the molecule interacts with other proteins, and reveals a potential new binding site distal to the IgE binding site. The results of this study point to new directions for the design of molecules to inhibit the interaction of Fc epsilon RI alpha with its natural ligand and thus to prevent a primary step in the allergic response.
- Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Twinbrook II, 12441 Parklawn Drive, Rockville, MD 20852, USA. garman@alpha.niaid.nih.gov
Organizational Affiliation: