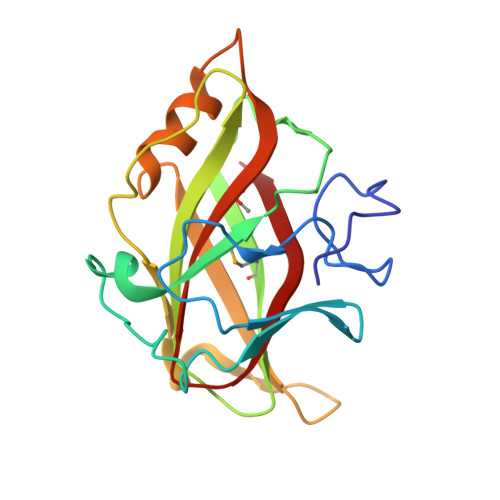
Recognition of cello-oligosaccharides by a family 17 carbohydrate-binding module: an X-ray crystallographic, thermodynamic and mutagenic study.
Notenboom, V., Boraston, A.B., Chiu, P., Freelove, A.C., Kilburn, D.G., Rose, D.R.(2001) J Mol Biology 314: 797-806
- PubMed: 11733998 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5153
- Primary Citation Related Structures:
1J83, 1J84 - PubMed Abstract:
The crystal structure of the Clostridium cellulovorans carbohydrate-binding module (CBM) belonging to family 17 has been solved to 1.7 A resolution by multiple anomalous dispersion methods. CBM17 binds to non-crystalline cellulose and soluble beta-1,4-glucans, with a minimal binding requirement of cellotriose and optimal affinity for cellohexaose. The crystal structure of CBM17 complexed with cellotetraose solved at 2.0 A resolution revealed that binding occurs in a cleft on the surface of the molecule involving two tryptophan residues and several charged amino acids. Thermodynamic binding studies and alanine scanning mutagenesis in combination with the cellotetraose complex structure allowed the mapping of the CBM17 binding cleft. In contrast to the binding groove characteristic of family 4 CBMs, family 17 CBMs appear to have a very shallow binding cleft that may be more accessible to cellulose chains in non-crystalline cellulose than the deeper binding clefts of family 4 CBMs. The structural differences in these two modules may reflect non-overlapping binding niches on cellulose surfaces.
- Protein Engineering Networks of Centres of Excellence, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: