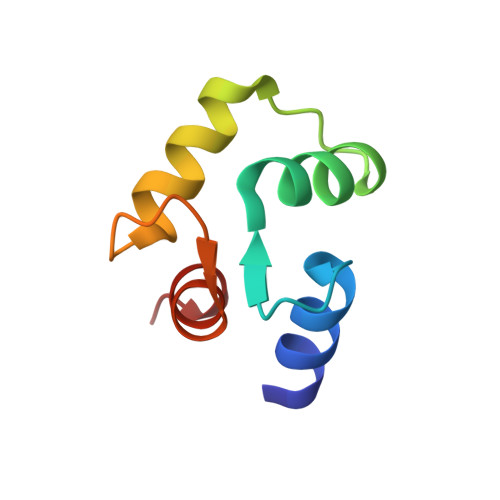
Solution structure of Ca(2+)-calmodulin reveals flexible hand-like properties of its domains.
Chou, J.J., Li, S., Klee, C.B., Bax, A.(2001) Nat Struct Biol 8: 990-997
- PubMed: 11685248 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1101-990
- Primary Citation Related Structures:
1J7O, 1J7P - PubMed Abstract:
The solution structure of Ca(2+)-ligated calmodulin is determined from residual dipolar couplings measured in a liquid crystalline medium and from a large number of heteronuclear J couplings for defining side chains. Although the C-terminal domain solution structure is similar to the X-ray crystal structure, the EF hands of the N-terminal domain are considerably less open. The substantial differences in interhelical angles correspond to negligible changes in short interproton distances and, therefore, cannot be identified by comparison of NOEs and X-ray data. NOE analysis, however, excludes a two-state equilibrium in which the closed apo conformation is partially populated in the Ca(2+)-ligated state. The difference between the crystal and solution structures of Ca(2+)-calmodulin indicates considerable backbone plasticity within the domains of calmodulin, which is key to their ability to bind a wide range of targets. In contrast, the vast majority of side chains making up the target binding surface are locked into the same chi(1) rotameric states as in complexes with target peptide.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892-0520, USA.
Organizational Affiliation: