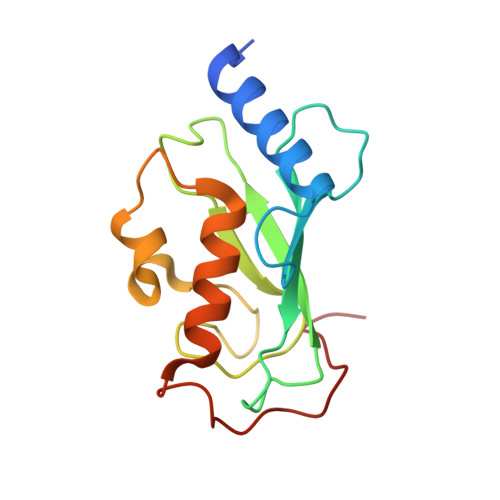
Crystal structure of the human ubiquitin conjugating enzyme complex, hMms2-hUbc13.
Moraes, T.F., Edwards, R.A., McKenna, S., Pastushok, L., Xiao, W., Glover, J.N., Ellison, M.J.(2001) Nat Struct Biol 8: 669-673
- PubMed: 11473255 Search on PubMed
- DOI: https://doi.org/10.1038/90373
- Primary Citation Related Structures:
1J74, 1J7D - PubMed Abstract:
The ubiquitin conjugating enzyme complex Mms2-Ubc13 plays a key role in post-replicative DNA repair in yeast and the NF-kappaB signal transduction pathway in humans. This complex assembles novel polyubiquitin chains onto yet uncharacterized protein targets. Here we report the crystal structure of a complex between hMms2 (Uev1) and hUbc13 at 1.85 A resolution and a structure of free hMms2 at 1.9 A resolution. These structures reveal that the hMms2 monomer undergoes a localized conformational change upon interaction with hUbc13. The nature of the interface provides a physical basis for the preference of Mms2 for Ubc13 as a partner over a variety of other structurally similar ubiquitin-conjugating enzymes. The structure of the hMms2-hUbc13 complex provides the conceptual foundation for understanding the mechanism of Lys 63 multiubiquitin chain assembly and for its interactions with the RING finger proteins Rad5 and Traf6.
- Department of Biochemistry, University of Alberta, Edmonton, Alberta, T6G-2H7, Canada.
Organizational Affiliation: