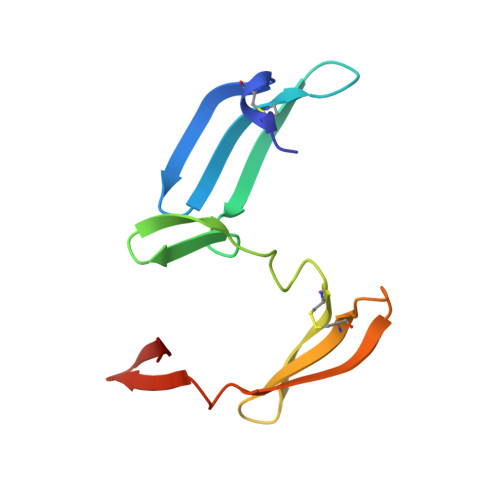
Using conjoined rigid body/torsion angle simulated annealing to determine the relative orientation of covalently linked protein domains from dipolar couplings.
Clore, G.M., Bewley, C.A.(2002) J Magn Reson (1969 154: 329-335
- PubMed: 11846592 Search on PubMed
- DOI: https://doi.org/10.1006/jmre.2001.2489
- Primary Citation Related Structures:
1J4V - PubMed Abstract:
A simple and robust method for determining the relative orientations of covalently linked protein domains using conjoined rigid body/torsion angle dynamics simulated annealing on the basis of residual dipolar couplings is presented. In this approach each domain is treated as a rigid body and the relevant degrees of conformational freedom are restricted to the backbone torsion angles (phi, psi) of the linker between the domains. By this means translational information afforded by the presence of an intact linker is preserved. We illustrate this approach using the domain-swapped dimer of the HIV-inactivating protein cyanovirin-N as an example.
- Laboratory of Chemical Physics, Building 5, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892-0510, USA. clore@speck.niddk.nih.gov
Organizational Affiliation: