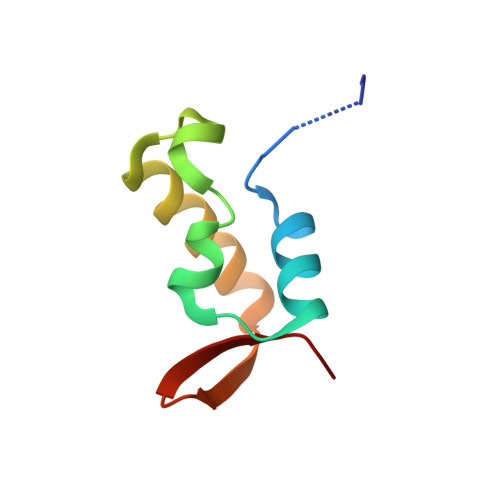
Molecular mechanism of recruitment of TFIIF- associating RNA polymerase C-terminal domain phosphatase (FCP1) by transcription factor IIF
Kamada, K., Roeder, R.G., Burley, S.K.(2003) Proc Natl Acad Sci U S A 100: 2296-2299
- PubMed: 12591941 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.262798199
- Primary Citation Related Structures:
1J2X - PubMed Abstract:
After mRNA transcription termination in eukaryotes, the hyperphosphorylated form of RNA polymerase II (pol II0) must be recycled by TFIIF-associating C-terminal domain phosphatase (FCP1), the phosphatase responsible for dephosphorylating the C-terminal domain of the largest polymerase subunit. Transcription factor (TF)-IIF stimulates the activity of FCP1, and the RNA polymerase II-associating protein 74 subunit of TFIIF forms a complex with FCP1 in both human and yeast. Here, we report a cocrystal structure of the winged-helix domain of human RNA polymerase II-associating protein 74 bound to the alpha-helical C terminus of human FCP1 (residues 944-961). These results illustrate the molecular mechanism by which TFIIF efficiently recruits FCP1 to the pol II transcription machinery for recycling of the polymerase.
- Laboratory of Molecular Biophysics and Howard Hughes Medical Institute, The Rockefeller University, 1230 York Avenue, New York, NY 10021, USA.
Organizational Affiliation: