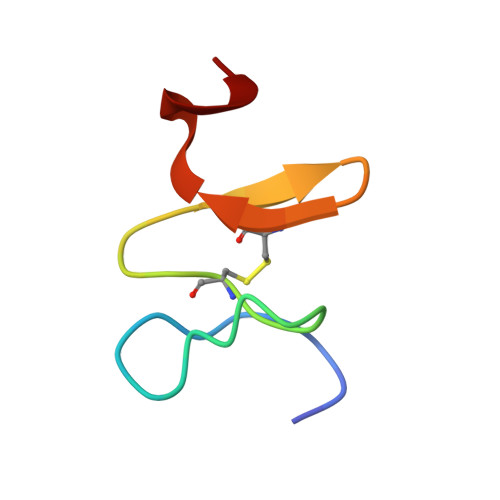
Structural Basis for New Pattern of Conserved Amino Acid Residues Related to Chitin-binding in the Antifungal Peptide from the Coconut Rhinoceros Beetle Oryctes rhinoceros
Hemmi, H., Ishibashi, J., Tomie, T., Yamakawa, M.(2003) J Biological Chem 278: 22820-22827
- PubMed: 12676931 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M301025200
- Primary Citation Related Structures:
1IYC - PubMed Abstract:
Scarabaecin isolated from hemolymph of the coconut rhinoceros beetle Oryctes rhinoceros is a 36-residue polypeptide that has antifungal activity. The solution structure of scarabaecin has been determined from twodimensional 1H NMR spectroscopic data and hybrid distance geometry-simulated annealing protocol calculation. Based on 492 interproton and 10 hydrogen-bonding distance restraints and 36 dihedral angle restraints, we obtained 20 structures. The average backbone root-mean-square deviation for residues 4-35 is 0.728 +/- 0.217 A from the mean structure. The solution structure consists of a two-stranded antiparallel beta-sheet connected by a type-I beta-turn after a short helical turn. All secondary structures and a conserved disulfide bond are located in the C-terminal half of the peptide, residues 18-36. Overall folding is stabilized by a combination of a disulfide bond, seven hydrogen bonds, and numerous hydrophobic interactions. The structural motif of the C-terminal half shares a significant tertiary structural similarity with chitin-binding domains of plant and invertebrate chitin-binding proteins, even though scarabaecin has no overall sequence similarity to other peptide/polypeptides including chitin-binding proteins. The length of its primary structure, the number of disulfide bonds, and the pattern of conserved functional residues binding to chitin in scarabaecin differ from those of chitin-binding proteins in other invertebrates and plants, suggesting that scarabaecin does not share a common ancestor with them. These results are thought to provide further strong experimental evidence to the hypothesis that chitin-binding proteins of invertebrates and plants are correlated by a convergent evolution process.
- National Food Research Institute, 2-1-12 Kannondai, Tsukuba, Ibaraki 305-8642, Japan. hemmi@nfri.affrc.go.jp
Organizational Affiliation: