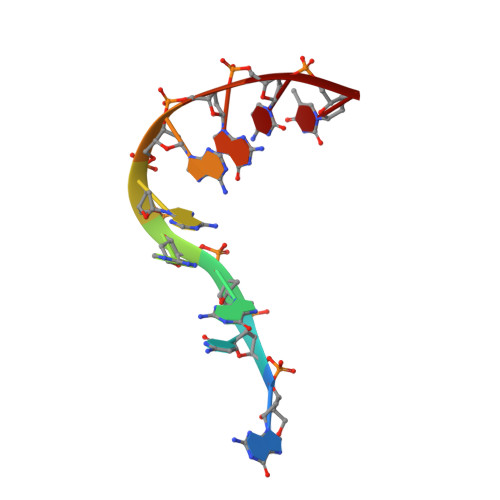
Crystal Structure of d(GCGAAAGCT) Containing a Parallel-stranded Duplex with Homo Base Pairs and an Anti-Parallel Duplex with Watson-Crick Base pairs
Sunami, T., Kondo, J., Kobuna, T., Hirao, I., Watanabe, K., Miura, K., Takenaka, A.(2002) Nucleic Acids Res 30: 5253-5260
- PubMed: 12466550 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkf639
- Primary Citation Related Structures:
1IXJ - PubMed Abstract:
A DNA fragment d(GCGAAAGCT), known to adopt a stable mini-hairpin structure in solution, has been crystallized in the space group I4(1)22 with the unit-cell dimensions a = b = 53.4 A and c = 54.0 A, and the crystal structure has been determined at 2.5 A resolution. The four nucleotide residues CGAA of the first half of the oligomer form a parallel duplex with another half through the homo base pairs, C2:C2+ (singly-protonated between the Watson- Crick sites), G3:G3 (between the minor groove sites), A4:A4 (between the major groove sites) and A5:A5 (between the Watson-Crick sites). The two strands remaining in the half of the parallel duplex are split away in different directions, and they pair in an anti-parallel B-form duplex with the second half extending from a neighboring parallel duplex, so that an infinite column is formed in a head-to-tail fashion along the c-axis. It seems that a hexa-ammine cobalt cation supports such a branched and bent conformation of the oligomer. One end of the parallel duplex is stacked on the corresponding end of the adjacent parallel duplex; between them, the guanine base of the first residue is stacked on the fourth ribose of another duplex.
- Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, Yokohama 226-8501, Japan.
Organizational Affiliation: