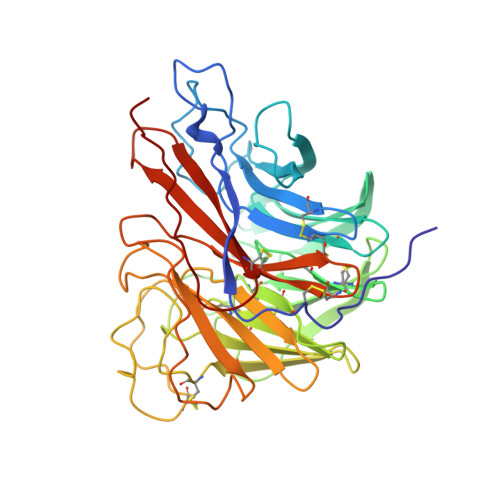
Structures of aromatic inhibitors of influenza virus neuraminidase.
Jedrzejas, M.J., Singh, S., Brouillette, W.J., Laver, W.G., Air, G.M., Luo, M.(1995) Biochemistry 34: 3144-3151
- PubMed: 7880809
- DOI: https://doi.org/10.1021/bi00010a003
- Primary Citation of Related Structures:
1IVB, 1IVC, 1IVD, 1IVE, 1IVF, 1IVG - PubMed Abstract:
Neuraminidase (NA), a surface glycoprotein of influenza virus, is a potential target for design of antiinfluenza agents. The crystal structure of influenza virus neuraminidase showed that in the active site 11 residues are universally conserved among all strains known so far. Several potent inhibitors based on the carbohydrate compound 2-deoxy-2,3-didehydro-D-N-acetylneuraminic acid (DANA) have been shown to bind to the conserved active site and to reduce virus infection in animals when administered by nasal spray. Inhibitors of this type are, however, rapidly excreted from physiological systems and may not be effective in order to provide long-time protection. A new class of specific NA inhibitors, which are benzoic acid derivatives, has been designed on the basis of the three-dimensional structure of the NA-DANA complex and modeling of derivatives of 4-(acetylamino)benzoic acid in the NA active site. Intermediates were synthesized and were shown to moderately inhibit the NA activity and to bind to the NA active site as predicted. These rudimentary inhibitors, 4-(acetylamino)-3-hydroxy-5-nitrobenzoic acid, 4-(acetylamino)-3-hydroxy-5-aminobenzoic acid, and 4-(acetylamino)-3-aminobenzoic acid, and their X-ray structures in complexes with N2 (A/Tokyo/3/67) and B/Lee/40 neuraminidases have been analyzed. The coordinates of such inhibitors complexed with NA were used as the starting model for further design of more potent benzoic acid inhibitors. Because the active site residues of NA are invariant, the designed aromatic inhibitors have the potential to become an antiviral drug against all strains of influenza virus.
- Department of Microbiology, University of Alabama at Birmingham 35294.
Organizational Affiliation: