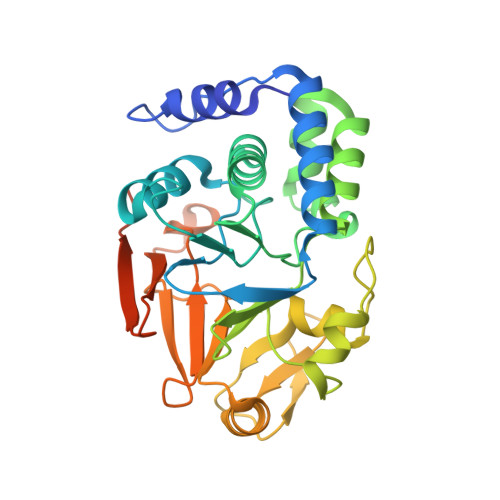
Crystal structure of the complex between calyculin A and the catalytic subunit of protein phosphatase 1.
Kita, A., Matsunaga, S., Takai, A., Kataiwa, H., Wakimoto, T., Fusetani, N., Isobe, M., Miki, K.(2002) Structure 10: 715-724
- PubMed: 12015153
- DOI: https://doi.org/10.1016/s0969-2126(02)00764-5
- Primary Citation of Related Structures:
1IT6 - PubMed Abstract:
The crystal structure of the catalytic subunit of the protein phosphatase 1 (PP1), PP1 gamma, in complex with a marine toxin, calyculin A, was determined at 2.0 A resolution. The metal binding site contains the phosphate group of calyculin A and forms a tight network via the hydrophilic interactions between PP1 and calyculin A. Calyculin A is located in two of the three grooves, namely, in the hydrophobic groove and the acidic groove on the molecular surface. This is the first observation to note that the inhibitor adopts not a pseudocyclic conformation but an extended conformation in order to form a complex with the protein. The amino acid terminus of calyculin A contributes, in a limited manner, to the binding to PP1 gamma, which is consistent with findings from the studies of dose-inhibition analysis.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: