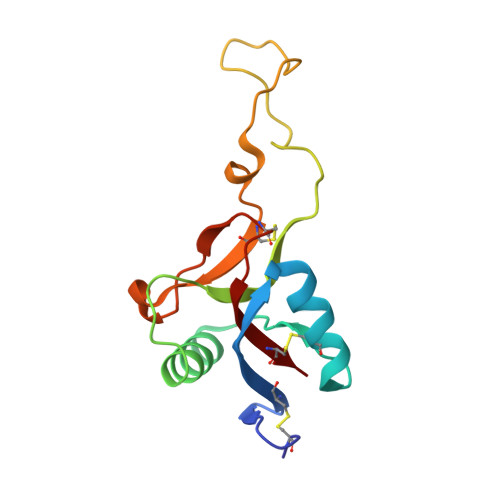
Crystal structure of an anticoagulant protein in complex with the Gla domain of factor X.
Mizuno, H., Fujimoto, Z., Atoda, H., Morita, T.(2001) Proc Natl Acad Sci U S A 98: 7230-7234
- PubMed: 11404471 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.131179698
- Primary Citation Related Structures:
1IOD - PubMed Abstract:
The gamma-carboxyglutamic acid (Gla) domain of blood coagulation factors is responsible for Ca2+-dependent phospholipid membrane binding. Factor X-binding protein (X-bp), an anticoagulant protein from snake venom, specifically binds to the Gla domain of factor X. The crystal structure of X-bp in complex with the Gla domain peptide of factor X at 2.3-A resolution showed that the anticoagulation is based on the fact that two patches of the Gla domain essential for membrane binding are buried in the complex formation. The Gla domain thus is expected to be a new target of anticoagulant drugs, and X-bp provides a basis for designing them. This structure also provides a membrane-bound model of factor X.
- Department of Biotechnology, National Institute of Agrobiological Resources, Tsukuba, Ibaraki 305-8602, Japan. mizuno@nias.affrc.go.jp
Organizational Affiliation: