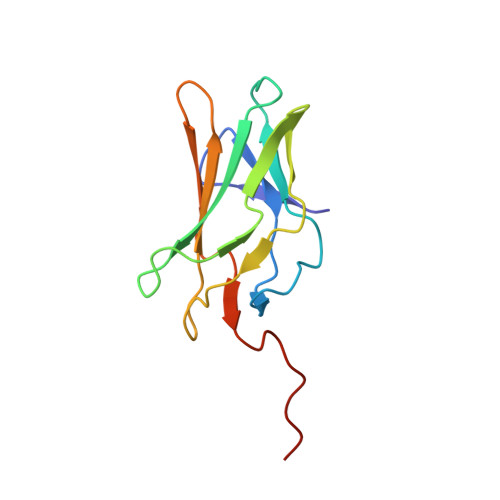
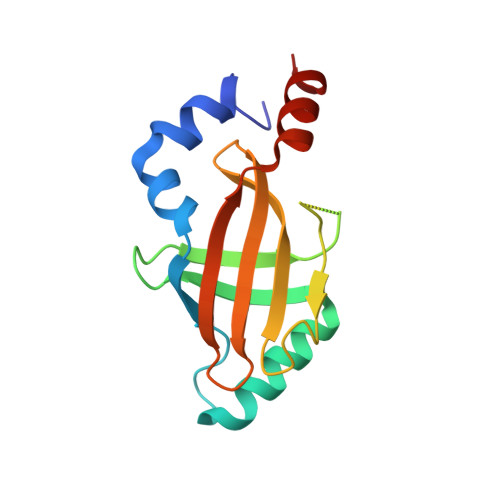
Structural analyses of DNA recognition by the AML1/Runx-1 Runt domain and its allosteric control by CBFbeta.
Tahirov, T.H., Inoue-Bungo, T., Morii, H., Fujikawa, A., Sasaki, M., Kimura, K., Shiina, M., Sato, K., Kumasaka, T., Yamamoto, M., Ishii, S., Ogata, K.(2001) Cell 104: 755-767
- PubMed: 11257229
- DOI: https://doi.org/10.1016/s0092-8674(01)00271-9
- Primary Citation of Related Structures:
1HJB, 1HJC, 1IO4 - PubMed Abstract:
The core binding factor (CBF) heterodimeric transcription factors comprised of AML/CBFA/PEBP2alpha/Runx and CBFbeta/PEBP2beta subunits are essential for differentiation of hematopoietic and bone cells, and their mutation is intimately related to the development of acute leukemias and cleidocranial dysplasia. Here, we present the crystal structures of the AML1/Runx-1/CBFalpha(Runt domain)-CBFbeta(core domain)-C/EBPbeta(bZip)-DNA, AML1/Runx-1/CBFalpha(Runt domain)-C/EBPbeta(bZip)-DNA, and AML1/Runx-1/CBFalpha(Runt domain)-DNA complexes. The hydrogen bonding network formed among CBFalpha(Runt domain) and CBFbeta, and CBFalpha(Runt domain) and DNA revealed the allosteric regulation mechanism of CBFalpha(Runt domain)-DNA binding by CBFbeta. The point mutations of CBFalpha related to the aforementioned diseases were also mapped and their effect on DNA binding is discussed.
- Kanagawa Academy of Science and Technology (KAST), Yokohama City University School of Medicine, 3-9 Fukuura, Yokohama 236-0004, Kanazawa-ku, Japan. tahir@med.yokohama-cu.ac.jp
Organizational Affiliation: