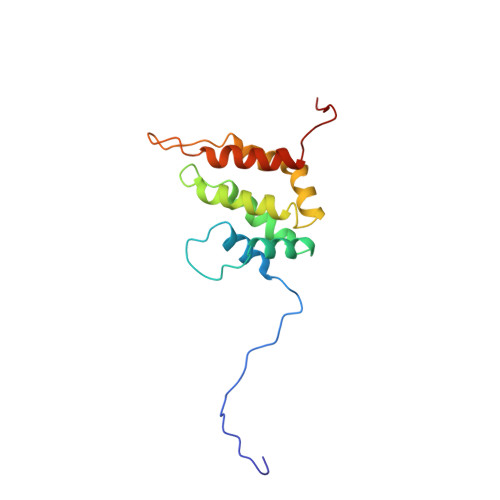
Solution structure of the epsin N-terminal homology (ENTH) domain of human epsin
Koshiba, S., Kigawa, T., Kikuchi, A., Yokoyama, S.(2001) J Struct Funct Genomics 2: 1-8
- PubMed: 12836669 Search on PubMed
- DOI: https://doi.org/10.1023/a:1011397007366
- Primary Citation Related Structures:
1INZ - PubMed Abstract:
Epsin is a protein that binds to the Eps15 homology (EH) domains, and is involved in clathrin-mediated endocytosis. The epsin N-terminal homology (ENTH) domain (about 140 amino acid residues) is well conserved in eukaryotes and is considered to be important for actin cytoskeleton organization in endocytosis. In this study, we have determined the solution structure of the ENTH domain (residues 1-144) of human epsin by multidimensional nuclear magnetic resonance spectroscopy. In the ENTH-domain structure, seven alpha-helices form a superhelical fold, consisting of two antiparallel two-helix HEAT motifs and one three-helix ARM motif, with a continuous hydrophobic core in the center. We conclude that the seven-helix superhelical fold defines the ENTH domain, and that the previously-reported eight-helix fold of a longer fragment of rat epsin 1 is divided into the authentic ENTH domain and a C-terminal flanking alpha-helix.
- Genomic Sciences Center, RIKEN Yokohama Institute, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan.
Organizational Affiliation: