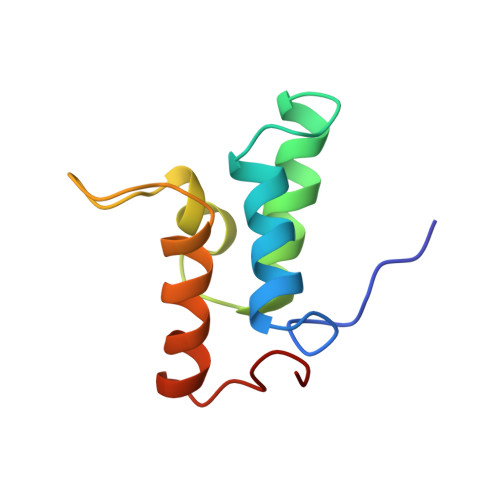
Three-dimensional solution structure and 13C nuclear magnetic resonance assignments of the colicin E9 immunity protein Im9.
Osborne, M.J., Breeze, A.L., Lian, L.Y., Reilly, A., James, R., Kleanthous, C., Moore, G.R.(1996) Biochemistry 35: 9505-9512
- PubMed: 8755730 Search on PubMed
- DOI: https://doi.org/10.1021/bi960401k
- Primary Citation Related Structures:
1IMP, 1IMQ - PubMed Abstract:
The 86-amino acid colicin E9 immunity protein (Im9), which inhibits the DNase activity of colicin E9, has been overexpressed in Escherichia coli and isotopically enriched with 15N and 13C. Using the 3D CBCANH and CBCA(CO)NH experiments, we have almost completely assigned the backbone 13C resonances and extended previously reported 15N/1H backbone assignments [Osborne et al. (1994), Biochemistry 33, 12347-12355]. Side chain assignments for almost all residues were made using the 3D 13C HCCH-TOCSY experiment allied to previous 1H assignments. Sixty solution structures of Im9 were determined using the DIANA program on the basis of 1210 distance restraints and 56 dihedral angle restraints. The 30 lowest-energy structures were then subjected to a slow-cooling simulated annealing protocol using XPLOR and the 21 lowest-energy structures, satisfying the geometric restraints chosen for further analysis. The Im9 structure is well-defined except for the termini and two solvent-exposed loops between residues 28-32 and 57-64. The average RMSD about the average structure of residues 4-84 was 0.94 A for all heavy atoms and 0.53 A for backbone C alpha, C = O, and N atoms. The Im9 fold is novel and can be considered a distorted antiparallel four-helix bundle, in which the third helix is rather short, being terminated close to its N-terminal end by a proline at its C-terminus. The structure fits in well with available kinetic and biochemical data concerning the interaction between Im9 and its target DNase. Important residues of Im9 that govern specificity are located on the molecular surface in a region rich in negatively charged groups, consistent with the proposed electrostatically steered association [Wallis et al. (1995a), Biochemistry 34, 13743-13750].
- School of Biological Science, University of East Anglia, Norwich, U.K.
Organizational Affiliation: