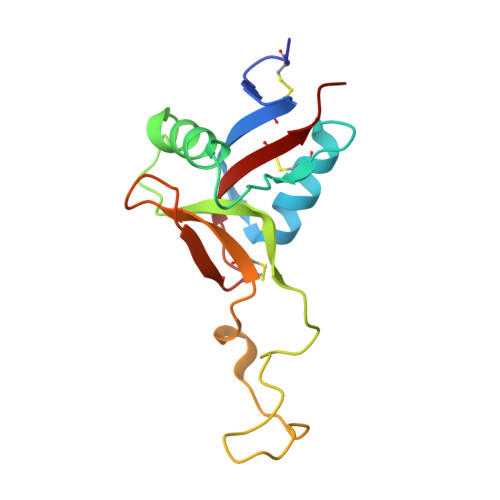
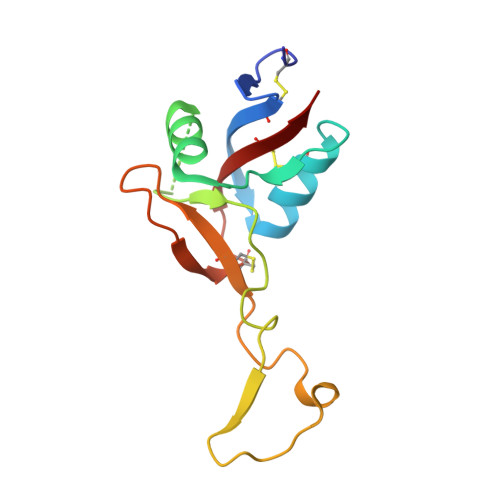
Structural basis of von Willebrand factor activation by the snake toxin botrocetin.
Fukuda, K., Doggett, T.A., Bankston, L.A., Cruz, M.A., Diacovo, T.G., Liddington, R.C.(2002) Structure 10: 943-950
- PubMed: 12121649 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00787-6
- Primary Citation Related Structures:
1IJB, 1IJK - PubMed Abstract:
The A1 domain of von Willebrand factor (vWF) mediates platelet adhesion to sites of vascular injury by binding to the platelet receptor glycoprotein Ib (GpIb), an interaction that is regulated by hydrodynamic shear forces. The GpIb binding surface of A1 is distinct from a regulatory region, suggesting that ligand binding is controlled allosterically. Here we report the crystal structures of the "gain-of-function" mutant A1 domain (I546V) and its complex with the exogenous activator botrocetin. We show that botrocetin switches the mutant A1 back toward the wild-type conformation, suggesting that affinity is enhanced by augmenting the GpIb binding surface rather than through allosteric control. Functional studies of platelet adhesion under flow further suggest that the activation mechanism is distinct from that of the gain-of-function mutation.
- The Burnham Institute, 10901 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: