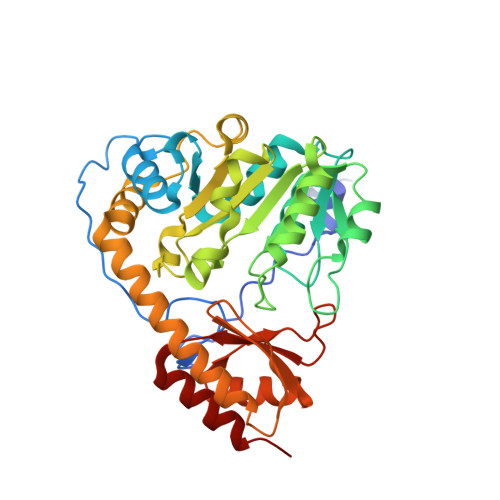
Crystal structure of histidinol phosphate aminotransferase (HisC) from Escherichia coli, and its covalent complex with pyridoxal-5'-phosphate and l-histidinol phosphate.
Sivaraman, J., Li, Y., Larocque, R., Schrag, J.D., Cygler, M., Matte, A.(2001) J Mol Biology 311: 761-776
- PubMed: 11518529 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4882
- Primary Citation Related Structures:
1FG3, 1FG7, 1IJI - PubMed Abstract:
The biosynthesis of histidine is a central metabolic process in organisms ranging from bacteria to yeast and plants. The seventh step in the synthesis of histidine within eubacteria is carried out by a pyridoxal-5'-phosphate (PLP)-dependent l-histidinol phosphate aminotransferase (HisC, EC 2.6.1.9). Here, we report the crystal structure of l-histidinol phosphate aminotransferase from Escherichia coli, as a complex with pyridoxamine-5'-phosphate (PMP) at 1.5 A resolution, as the internal aldimine with PLP, and in a covalent, tetrahedral complex consisting of PLP and l-histidinol phosphate attached to Lys214, both at 2.2 A resolution. This covalent complex resembles, in structural terms, the gem-diamine intermediate that is formed transiently during conversion of the internal to external aldimine.HisC is a dimeric enzyme with a mass of approximately 80 kDa. Like most PLP-dependent enzymes, each HisC monomer consists of two domains, a larger PLP-binding domain having an alpha/beta/alpha topology, and a smaller domain. An N-terminal arm contributes to the dimerization of the two monomers. The PLP-binding domain of HisC shows weak sequence similarity, but significant structural similarity with the PLP-binding domains of a number of PLP-dependent enzymes. Residues that interact with the PLP cofactor, including Tyr55, Asn157, Asp184, Tyr187, Ser213, Lys214 and Arg222, are conserved in the family of aspartate, tyrosine and histidinol phosphate aminotransferases. The imidazole ring of l-histidinol phosphate is bound, in part, through a hydrogen bond with Tyr110, a residue that is substituted by Phe in the broad substrate specific HisC enzymes from Zymomonas mobilis and Bacillus subtilis. Comparison of the structures of the HisC internal aldimine, the PMP complex and the HisC l-histidinol phosphate complex reveal minimal changes in protein or ligand structure. Proton transfer, required for conversion of the gem-diamine to the external aldimine, does not appear to be limited by the distance between substrate and lysine amino groups. We propose that the tetrahedral complex has resulted from non-productive binding of l-histidinol phosphate soaked into the HisC crystals, resulting in its inability to be converted to the external aldimine at the HisC active site.
- Biotechnology Research Institute, 6100 Royalmount Ave., Montreal, H4P 2R2 and, Canada.
Organizational Affiliation: