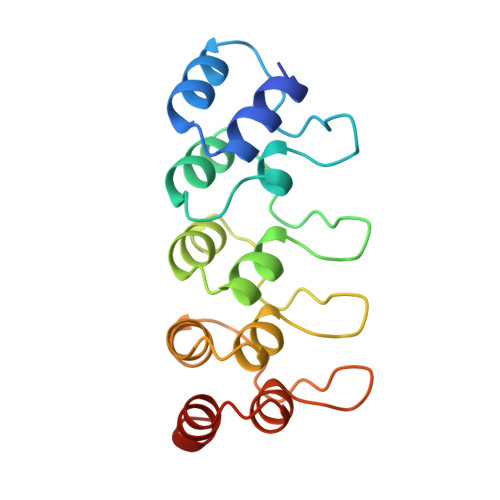
Crystal structure of the CDK4/6 inhibitory protein p18INK4c provides insights into ankyrin-like repeat structure/function and tumor-derived p16INK4 mutations.
Venkataramani, R., Swaminathan, K., Marmorstein, R.(1998) Nat Struct Biol 5: 74-81
- PubMed: 9437433 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0198-74
- Primary Citation Related Structures:
1IHB - PubMed Abstract:
p18INK4c is a member of a family of INK4 proteins that function to arrest the G1 to S cell cycle transition by inhibiting the activity of the cyclin-dependent kinases 4 and 6. The X-ray crystal structure of the human p18INK4c protein to a resolution of 1.95 A reveals an elongated molecule comprised of five contiguous 32- or 33-residue ankyrin-like repeat units. Each ankyrin-like repeat contains a beta-strand helix-turn-helix extended strand beta-strand motif that associates with neighboring motifs through beta-sheet, and helical bundle interactions. Conserved ankyrin-like repeat residues function to facilitate the ankyrin repeat fold and the tertiary interactions between neighboring repeat units. A large percentage of residues that are conserved among INK4 proteins and that map to positions of tumor-derived p16INK4 mutations play important roles in protein stability. A subset of these residues suggest an INK4 binding surface for the cyclin-dependent kinases 4 and 6. This surface is centered around a region that shows structural features uncharacteristic of ankyrin-like repeat units.
- Wistar Institute, Philadelphia, Pennsylvania, USA.
Organizational Affiliation: