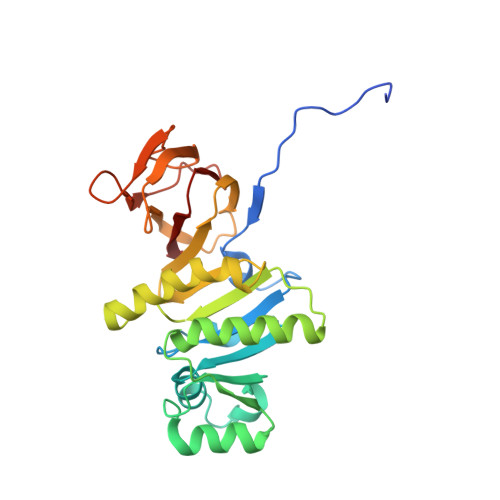
Crystal structure of thiamin pyrophosphokinase.
Timm, D.E., Liu, J., Baker, L.J., Harris, R.A.(2001) J Mol Biology 310: 195-204
- PubMed: 11419946
- DOI: https://doi.org/10.1006/jmbi.2001.4727
- Primary Citation Related Structures:
1IG3 - PubMed Abstract:
Thiamin pyrophosphate (TPP) is a coenzyme derived from vitamin B1 (thiamin). TPP synthesis in eukaryotes requires thiamin pyrophosphokinase (TPK), which catalyzes the transfer of a pyrophosphate group from ATP to thiamin. TPP is essential for central metabolic processes, including the formation of acetyl CoA from glucose and the Krebs cycle. Deficiencies in human thiamin metabolism result in beriberi and Wernicke encephalopathy. The crystal structure of mouse TPK was determined by multiwavelength anomalous diffraction at 2.4 A resolution, and the structure of TPK complexed with thiamin has been refined at 1.9 A resolution. The TPK polypeptide folds as an alpha/beta-domain and a beta-sandwich domain, which share a central ten-stranded mixed beta-sheet. TPK subunits associate as a dimer, and thiamin is bound in the dimer interface. Despite lacking apparent sequence homology with other proteins, the alpha/beta-domain resembles the Rossman fold and is similar to other kinase structures, including another pyrophosphokinase and a thiamin biosynthetic enzyme. Comparison of mouse and yeast TPK structures reveals differences that could be exploited in developing species-specific inhibitors of potential use as antimicrobial agents.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, IN 46202, USA. dtimm@iupui.edu
Organizational Affiliation: