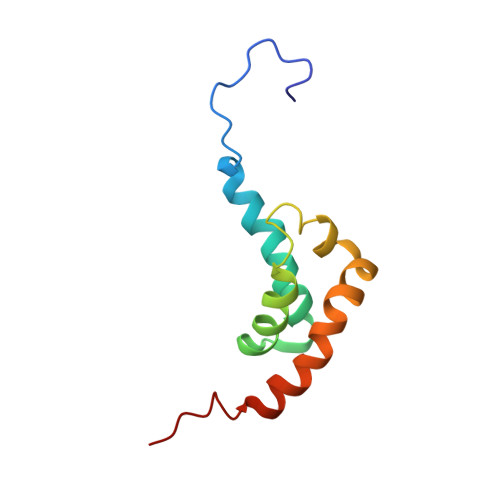
Solution structure of the orphan PABC domain from Saccharomyces cerevisiae poly(A)-binding protein.
Kozlov, G., Siddiqui, N., Coillet-Matillon, S., Trempe, J.F., Ekiel, I., Sprules, T., Gehring, K.(2002) J Biological Chem 277: 22822-22828
- PubMed: 11940585 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M201230200
- Primary Citation Related Structures:
1IFW - PubMed Abstract:
We have determined the solution structure of the PABC domain from Saccharomyces cerevisiae Pab1p and mapped its peptide-binding site. PABC domains are peptide binding domains found in poly(A)-binding proteins (PABP) and are a subset of HECT-family E3 ubiquitin ligases (also known as hyperplastic discs proteins (HYDs)). In mammals, the PABC domain of PABP functions to recruit several different translation factors to the mRNA poly(A) tail. PABC domains are highly conserved, with high specificity for peptide sequences of roughly 12 residues with conserved alanine, phenylalanine, and proline residues at positions 7, 10, and 12. Compared with human PABP, the yeast PABC domain is missing the first alpha helix, contains two extra amino acids between helices 2 and 3, and has a strongly bent C-terminal helix. These give rise to unique peptide binding specificity wherein yeast PABC binds peptides from Paip2 and RF3 but not Paip1. Mapping of the peptide-binding site reveals that the bend in the C-terminal helix disrupts binding interactions with the N terminus of peptide ligands and leads to greatly reduced binding affinity for the peptides tested. No high affinity or natural binding partners from S. cerevisiae could be identified by sequence analysis of known PABC ligands. Comparison of the three known PABC structures shows that the features responsible for peptide binding are highly conserved and responsible for the distinct but overlapping binding specificities.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6, Canada.
Organizational Affiliation: