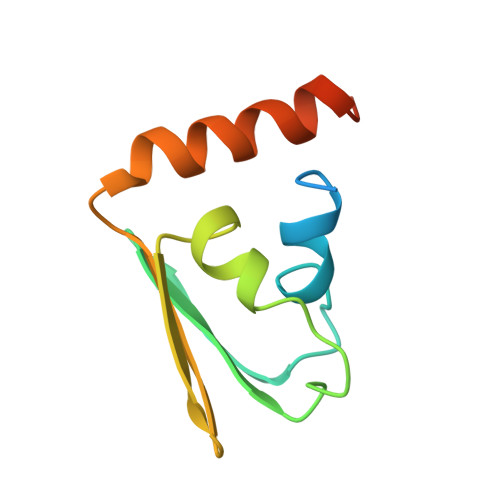
Solution structure of the IIB domain of the glucose transporter of Escherichia coli.
Eberstadt, M., Grdadolnik, S.G., Gemmecker, G., Kessler, H., Buhr, A., Erni, B.(1996) Biochemistry 35: 11286-11292
- PubMed: 8784182 Search on PubMed
- DOI: https://doi.org/10.1021/bi960492l
- Primary Citation Related Structures:
1IBA - PubMed Abstract:
The structure of the IIBGlc domain of the Escherichia coli transporter for glucose was determined by multidimensional heteronuclear NMR. The glucose transporter (IICBGlc) belongs to the bacterial phosphotransferase system. It mediates uptake with concomittant phosphorylation of glucose. The N-terminal IICGlc domain spans the membrane, the C-terminal IIBGlc domain (residues 386-477) contains the phosphorylation site, Cys421. The structure of the subclonal IIB domain was determined based on 927 conformational constraints, including 744 NOE derived upper bounds, 43 constraints of ranges of dihedral angles based on measurements of vicinal coupling constants, and 70 upper and lower bound constraints associated with 35 hydrogen bonds. The distance geometry interpretation of the NMR data is based on the previously published sequence-specific 1H, 15N, and 13C resonance assignments [Golic Grdadolnik et al. (1994) Eur. J. Biochem. 219, 945-952]. The sequence of the secondary structure elements of IIB is alpha 1 beta 1 beta 2 alpha 2 beta 3 beta 4 alpha 3. The basic fold consists of a split alpha/beta-sandwich composed of an antiparallel sheet with strand order beta 1 beta 2 beta 4 beta 3 and three alpha-helices superimposed onto one side of the sheet. The hydrophobic helix alpha 1 is packed against helices alpha 2, alpha 3, and the beta-sheet. The phosphorylation site (Cys421) is at the end of beta 1 on the solvent-exposed face of the sheet surrounded by Asp419, Thr423 Arg424, Arg426, and Gln456 which are invariant in 15 homologous IIB domains from other PTS transporters.
- Institut für Organische Chemie und Biochemie, Technische Universität München, Germany.
Organizational Affiliation: