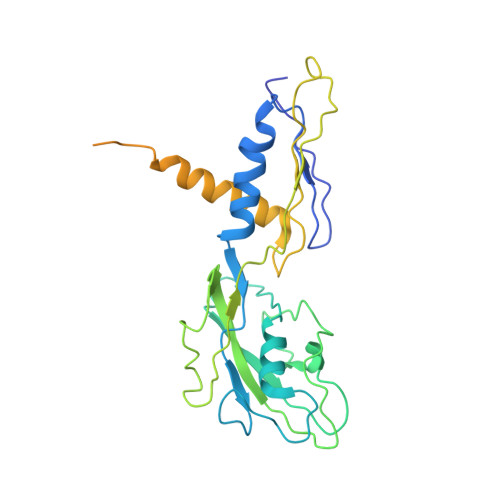
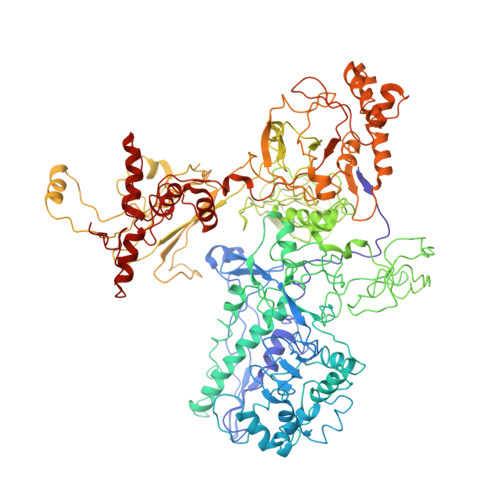
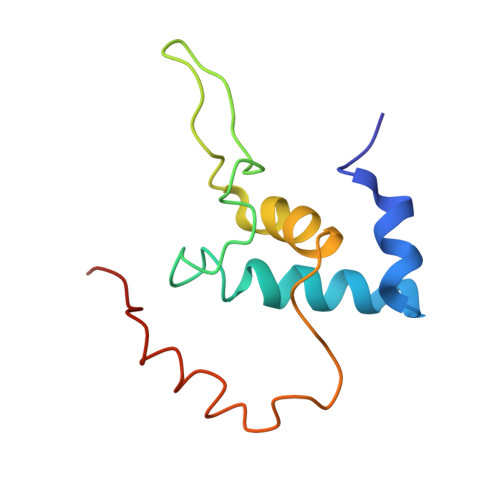
Structural mechanism for rifampicin inhibition of bacterial rna polymerase.
Campbell, E.A., Korzheva, N., Mustaev, A., Murakami, K., Nair, S., Goldfarb, A., Darst, S.A.(2001) Cell 104: 901-912
- PubMed: 11290327 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(01)00286-0
- Primary Citation Related Structures:
1I6V - PubMed Abstract:
Rifampicin (Rif) is one of the most potent and broad spectrum antibiotics against bacterial pathogens and is a key component of anti-tuberculosis therapy, stemming from its inhibition of the bacterial RNA polymerase (RNAP). We determined the crystal structure of Thermus aquaticus core RNAP complexed with Rif. The inhibitor binds in a pocket of the RNAP beta subunit deep within the DNA/RNA channel, but more than 12 A away from the active site. The structure, combined with biochemical results, explains the effects of Rif on RNAP function and indicates that the inhibitor acts by directly blocking the path of the elongating RNA when the transcript becomes 2 to 3 nt in length.
- The Rockefeller University, 1230 York Avenue, New York, NY 10021, USA.
Organizational Affiliation: