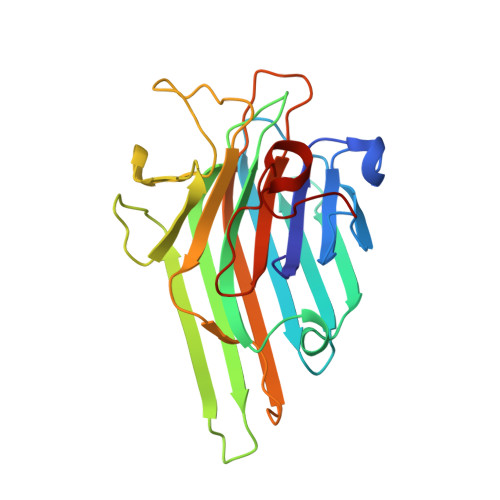
The 1.2 A resolution structure of the Con A-dimannose complex.
Sanders, D.A., Moothoo, D.N., Raftery, J., Howard, A.J., Helliwell, J.R., Naismith, J.H.(2001) J Mol Biology 310: 875-884
- PubMed: 11453694 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4806
- Primary Citation Related Structures:
1I3H - PubMed Abstract:
The complex between concanavalin A (Con A) and alpha1-2 mannobiose (mannose alpha1-2 mannose) has been refined to 1.2 A resolution. This is the highest resolution structure reported for any sugar-lectin complex. As the native structure of Con A to 0.94 A resolution is already in the database, this gives us a unique opportunity to examine sugar-protein binding at high resolution. These data have allowed us to model a number of hydrogen atoms involved in the binding of the sugar to Con A, using the difference density map to place the hydrogen atoms. This map reveals the presence of the protonated form of Asp208 involved in binding. Asp208 is not protonated in the 0.94 A native structure. Our results clearly show that this residue is protonated and hydrogen bonds to the sugar. The structure accounts for the higher affinity of the alpha1-2 linked sugar when compared to other disaccharides. This structure identifies different interactions to those predicted by previous modelling studies. We believe that the additional data presented here will enable significant improvements to be made to the sugar-protein modelling algorithms.
- Biomolecular Sciences, The University, St. Andrews, KY16 9ST, Scotland.
Organizational Affiliation: