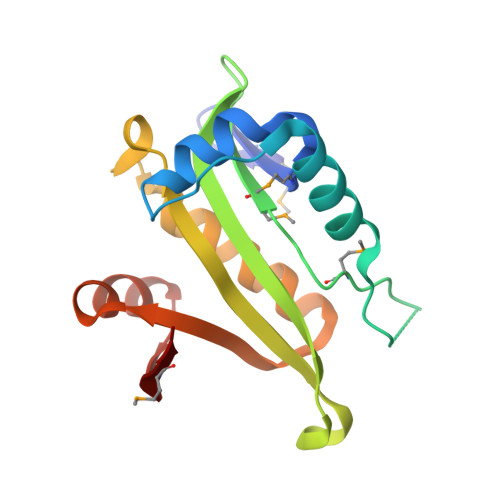
The crystal structures of Apo and complexed Saccharomyces cerevisiae GNA1 shed light on the catalytic mechanism of an amino-sugar N-acetyltransferase.
Peneff, C., Mengin-Lecreulx, D., Bourne, Y.(2001) J Biological Chem 276: 16328-16334
- PubMed: 11278591 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M009988200
- Primary Citation Related Structures:
1I12, 1I1D, 1I21 - PubMed Abstract:
The yeast enzymes involved in UDP-GlcNAc biosynthesis are potential targets for antifungal agents. GNA1, a novel member of the Gcn5-related N-acetyltransferase (GNAT) superfamily, participates in UDP-GlcNAc biosynthesis by catalyzing the formation of GlcNAc6P from AcCoA and GlcN6P. We have solved three crystal structures corresponding to the apo Saccharomyces cerevisiae GNA1, the GNA1-AcCoA, and the GNA1-CoA-GlcNAc6P complexes and have refined them to 2.4, 1.3, and 1.8 A resolution, respectively. These structures not only reveal a stable, beta-intertwined, dimeric assembly with the GlcNAc6P binding site located at the dimer interface but also shed light on the catalytic machinery of GNA1 at an atomic level. Hence, they broaden our understanding of structural features required for GNAT activity, provide structural details for related aminoglycoside N-acetyltransferases, and highlight the adaptability of the GNAT superfamily members to acquire various specificities.
- UMR 6098 CNRS, 31 chemin Joseph Aiguier, 13402 Marseille Cedex 20, France and UMR 8619 CNRS, Université Paris-Sud, Bâtiment 430, 91405 Orsay Cedex, France.
Organizational Affiliation: