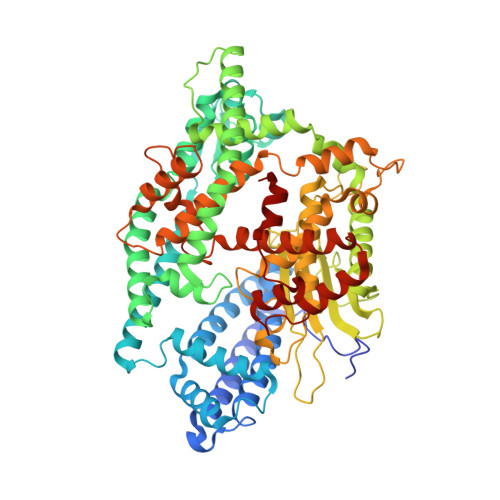
Structure of neurolysin reveals a deep channel that limits substrate access.
Brown, C.K., Madauss, K., Lian, W., Beck, M.R., Tolbert, W.D., Rodgers, D.W.(2001) Proc Natl Acad Sci U S A 98: 3127-3132
- PubMed: 11248043
- DOI: https://doi.org/10.1073/pnas.051633198
- Primary Citation Related Structures:
1I1I - PubMed Abstract:
The zinc metallopeptidase neurolysin is shown by x-ray crystallography to have large structural elements erected over the active site region that allow substrate access only through a deep narrow channel. This architecture accounts for specialization of this neuropeptidase to small bioactive peptide substrates without bulky secondary and tertiary structures. In addition, modeling studies indicate that the length of a substrate N-terminal to the site of hydrolysis is restricted to approximately 10 residues by the limited size of the active site cavity. Some structural elements of neurolysin, including a five-stranded beta-sheet and the two active site helices, are conserved with other metallopeptidases. The connecting loop regions of these elements, however, are much extended in neurolysin, and they, together with other open coil elements, line the active site cavity. These potentially flexible elements may account for the ability of the enzyme to cleave a variety of sequences.
- Department of Molecular and Cellular Biochemistry and Center for Structural Biology, University of Kentucky, Lexington, KY 40536, USA.
Organizational Affiliation: