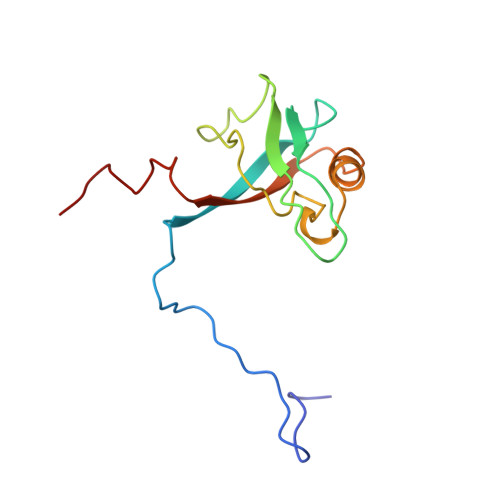
Structure of interleukin 16 resembles a PDZ domain with an occluded peptide binding site.
Muhlhahn, P., Zweckstetter, M., Georgescu, J., Ciosto, C., Renner, C., Lanzendorfer, M., Lang, K., Ambrosius, D., Baier, M., Kurth, R., Holak, T.A.(1998) Nat Struct Biol 5: 682-686
- PubMed: 9699630 Search on PubMed
- DOI: https://doi.org/10.1038/1376
- Primary Citation Related Structures:
1I16 - PubMed Abstract:
The structure of a folded core of IL-16 is similar to that of intracellular protein modules called PDZ domains. IL-16 is thus the first extracellular protein found to have a PDZ-like fold. However, it does not exhibit normal peptide binding properties of PDZ domains. This is due to alterations of the structure at the 'PDZ-like binding site' of IL-16 (the GLGF cleft): the GLGF cleft of IL-16 is much smaller than those of PDZ-domains and is additionally blocked with a tryptophan side chain at its center. Our experiments indicate also that IL-16 nonspecifically aggregates in solution; but formation of a homo-tetrameric protein is not required, in contrast to previous suggestions, for its chemo-attractant activity.
- Max Planck Institute for Biochemistry, Martinsried, F.R.G., Germany.
Organizational Affiliation: