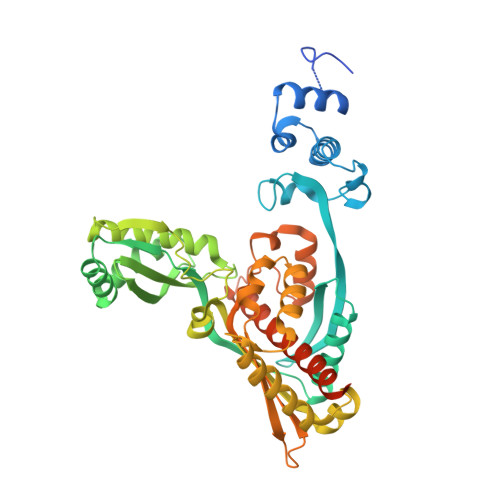
Structural mechanism for statin inhibition of HMG-CoA reductase.
Istvan, E.S., Deisenhofer, J.(2001) Science 292: 1160-1164
- PubMed: 11349148
- DOI: https://doi.org/10.1126/science.1059344
- Primary Citation of Related Structures:
1HW8, 1HW9, 1HWI, 1HWJ, 1HWK, 1HWL - PubMed Abstract:
HMG-CoA (3-hydroxy-3-methylglutaryl-coenzyme A) reductase (HMGR) catalyzes the committed step in cholesterol biosynthesis. Statins are HMGR inhibitors with inhibition constant values in the nanomolar range that effectively lower serum cholesterol levels and are widely prescribed in the treatment of hypercholesterolemia. We have determined structures of the catalytic portion of human HMGR complexed with six different statins. The statins occupy a portion of the binding site of HMG-CoA, thus blocking access of this substrate to the active site. Near the carboxyl terminus of HMGR, several catalytically relevant residues are disordered in the enzyme-statin complexes. If these residues were not flexible, they would sterically hinder statin binding.
- Department of Biochemistry, Howard Hughes Medical Institute, University of Texas Southwestern Medical Center at Dallas, TX 75390-9050, USA.
Organizational Affiliation: