Nonpeptide cyclic cyanoguanidines as HIV-1 protease inhibitors: synthesis, structure-activity relationships, and X-ray crystal structure studies.
Jadhav, P.K., Woerner, F.J., Lam, P.Y., Hodge, C.N., Eyermann, C.J., Man, H.W., Daneker, W.F., Bacheler, L.T., Rayner, M.M., Meek, J.L., Erickson-Viitanen, S., Jackson, D.A., Calabrese, J.C., Schadt, M., Chang, C.H.(1998) J Med Chem 41: 1446-1455
- PubMed: 9554878 Search on PubMed
- DOI: https://doi.org/10.1021/jm970524i
- Primary Citation Related Structures:
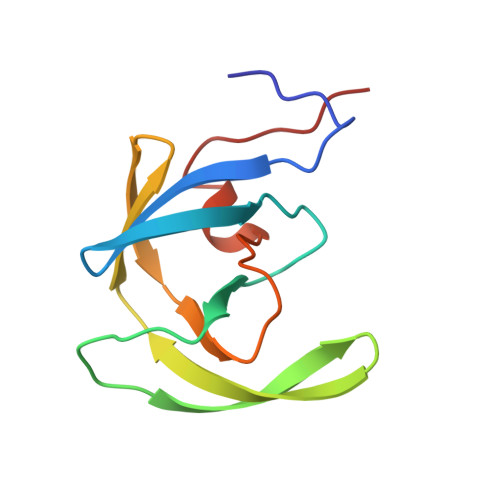
1HVH - PubMed Abstract:
Comparison of the high-resolution X-ray structures of the native HIV-1 protease and its complexes with the inhibitors suggested that the enzyme flaps are flexible. The movement at the tip of the flaps could be as large as 7 A. On the basis of this observation, cyclic cyanoguanidines have been designed, synthesized, and evaluated as HIV-1 protease (PR) inhibitors. Cyclic cyanoguanidines were found to be very potent inhibitors of HIV-1 protease. The choice of cyclic cyanoguanidines over cyclic guanidines was based on the reduced basicity of the former. X-ray structure studies of the HIV PR complex with cyclic cyanoguanidine demonstrated that in analogy to cyclic urea, cyclic cyanoguanidines also displace the unique structural water molecule. The structure-activity relationship of the cyclic cyanoguanidines is compared with that of the corresponding cyclic urea analogues. The differences in binding constants of the two series of compounds have been rationalized using high-resolution X-ray structure information.
- Chemical and Physical Sciences Department, The DuPont Merck Pharmaceutical Company, Experimental Station, P.O. Box 80500, Wilmington, Delaware 19880-0500, USA. Prabhakar.K.Jadhav@dupontmerck.com
Organizational Affiliation: