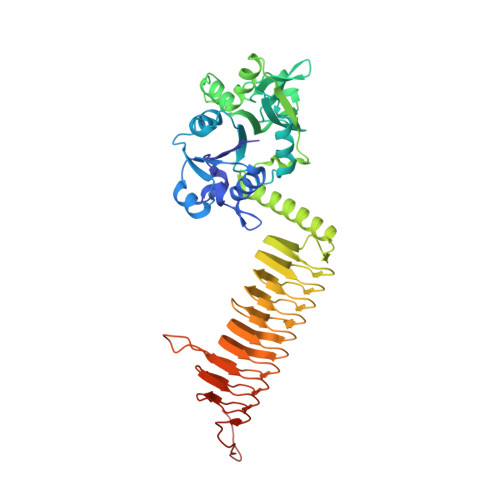
Structure of the Escherichia coli GlmU pyrophosphorylase and acetyltransferase active sites.
Olsen, L.R., Roderick, S.L.(2001) Biochemistry 40: 1913-1921
- PubMed: 11329257 Search on PubMed
- DOI: https://doi.org/10.1021/bi002503n
- Primary Citation Related Structures:
1HV9 - PubMed Abstract:
N-Acetylglucosamine-1-PO(4) uridyltransferase (GlmU) is a trimeric bifunctional enzyme that catalyzes the last two sequential reactions in the de novo biosynthetic pathway for UDP-GlcNAc. The X-ray crystal structure of Escherichia coli GlmU in complex with UDP-GlcNAc and CoA has been determined to 2.1 A resolution and reveals a two-domain architecture that is responsible for these two reactions. The C-terminal domain is responsible for the CoA-dependent acetylation of Glc-1-PO(4) to GlcNAc-1-PO(4) and displays the longest left-handed parallel beta-helix observed to date. The acetyltransferase active site defined by the binding site for CoA makes use of residues from all three subunits and is positioned beneath an open cavity large enough to accommodate the Glc-1-PO(4) acetyl acceptor. The N-terminal domain catalyzes uridyl transfer from UTP to GlcNAc-1-PO(4) to form the final products UDP-GlcNAc and pyrophosphate. This domain is composed of a central seven-stranded beta-sheet surrounded by six alpha-helices in a Rossmann fold-like topology. A Co(2+) ion binds to just one of the two independent pyrophosphorylase active sites present in the crystals studied here, each of which nonetheless binds UDP-GlcNAc. The conformational changes of the enzyme and sugar nucleotide that accompany metal binding may provide a window into the structural dynamics that accompany catalysis.
- Department of Biochemistry, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, New York 10461, USA.
Organizational Affiliation: