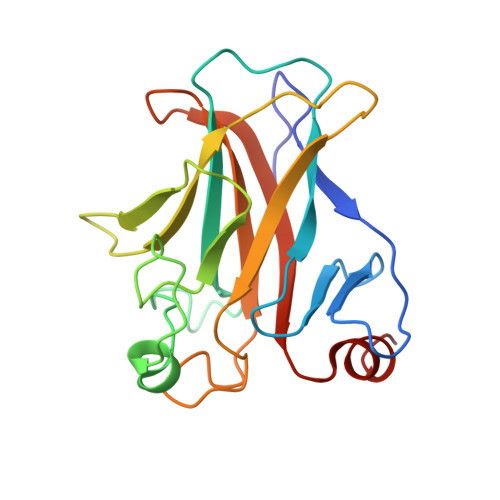
Crystal structure of the mouse p53 core DNA-binding domain at 2.7 A resolution.
Zhao, K., Chai, X., Johnston, K., Clements, A., Marmorstein, R.(2001) J Biological Chem 276: 12120-12127
- PubMed: 11152481 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M011644200
- Primary Citation Related Structures:
1HU8 - PubMed Abstract:
The p53 tumor suppressor is a sequence-specific DNA-binding protein that activates transcription in response to DNA damage to promote cell cycle arrest or apoptosis. The p53 protein functions in a tetrameric form in vivo and contains four domains including an N-terminal transcriptional activation domain, a C-terminal regulatory domain, a tetramerization domain, and a central core DNA-binding domain that is the site of the majority of tumor-derived mutations. Here we report the 2.7-A crystal structure of the mouse p53 core domain. Like the human p53 core domain in complex with DNA, the mouse p53 core domain adopts an immunoglobulin-like beta sandwich architecture with a series of loops and short helices at opposite ends of the beta sandwich. Comparison of the DNA-bound and DNA-free p53 core domains reveals that while the central beta sandwich architecture remains largely unchanged, a loop region important for DNA binding undergoes significant rearrangement. Although this loop region mediates major groove DNA contacts in the DNA-bound structure, it adopts a conformation that is incompatible with DNA binding in the DNA-free structure. Interestingly, crystals of the DNA-free core domain contain a noncrystallographic trimer with three nearly identical subunit-subunit (dimer) contacts. These dimer contacts align the p53 core domains in a way that is incompatible with simultaneous DNA binding by both protomers of the dimer. Surprisingly, similar dimer contacts are observed in crystals of the human p53 core domain with DNA in which only one of the three p53 protomers in the asymmetric unit cell is specifically bound to DNA. We propose that the p53 core domain dimer that is seen in the crystals described here represents a physiologically relevant inactive form of p53 that must undergo structural rearrangement for sequence-specific DNA binding.
- The Wistar Institute and the Department of Chemistry, University of Pennsylvania, 19104, USA.
Organizational Affiliation: