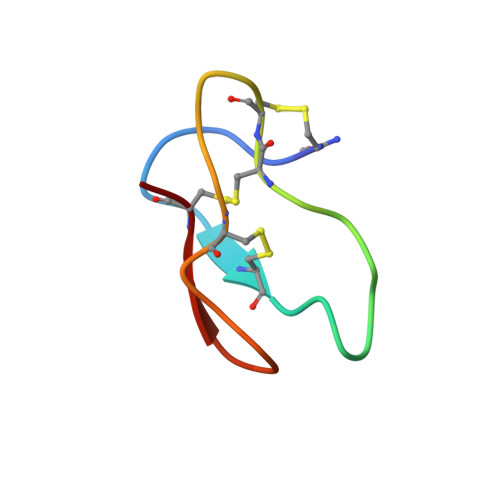
Solution structure of the main alpha-amylase inhibitor from amaranth seeds.
Martins, J.C., Enassar, M., Willem, R., Wieruzeski, J.M., Lippens, G., Wodak, S.J.(2001) Eur J Biochem 268: 2379-2389
- PubMed: 11298757
- DOI: https://doi.org/10.1046/j.1432-1327.2001.02118.x
- Primary Citation Related Structures:
1HTX - PubMed Abstract:
The most abundant alpha-amylase inhibitor (AAI) present in the seeds of Amaranthus hypochondriacus, a variety of the Mexican crop plant amaranth, is the smallest polypeptide (32 residues) known to inhibit alpha-amylase activity of insect larvae while leaving that of mammals unaffected. In solution, 1H NMR reveals that AAI isolated from amaranth seeds adopts a major trans (70%) and minor cis (30%) conformation, resulting from slow cis-trans isomerization of the Val15-Pro16 peptide bond. Both solution structures have been determined using 2D 1H-NMR spectroscopy and XPLOR followed by restrained energy refinement in the consistent-valence force field. For the major isomer, a total of 563 distance restraints, including 55 medium-range and 173 long-range ones, were available from the NOESY spectra. This rather large number of constraints from a protein of such a small size results from a compact fold, imposed through three disulfide bridges arranged in a cysteine-knot motif. The structure of the minor cis isomer has also been determined using a smaller constraint set. It reveals a different backbone conformation in the Pro10-Pro20 segment, while preserving the overall global fold. The energy-refined ensemble of the major isomer, consisting of 20 low-energy conformers with an average backbone rmsd of 0.29 +/- 0.19 A and no violations larger than 0.4 A, represents a considerable improvement in precision over a previously reported and independently performed calculation on AAI obtained through solid-phase synthesis, which was determined with only half the number of medium-range and long-range restraints reported here, and featured the trans isomer only. The resulting differences in ensemble precision have been quantified locally and globally, indicating that, for regions of the backbone and a good fraction of the side chains, the conformation is better defined in the new solution structure. Structural comparison of the solution structure with the X-ray structure of the inhibitor when bound to its alpha-amylase target in Tenebrio molitor shows that the backbone conformation is only slightly adjusted on complexation, while that of the side chains involved in protein-protein contacts is similar to those present in solution. Therefore, the overall conformation of AAI appears to be predisposed to binding to its target alpha-amylase, confirming the view that it acts as a lid on top of the alpha-amylase active site.
- High Resolution NMR Centrum (HNMR), Vrije Universiteit Brussel, Belgium. jose@hnmr.vub.ac.be
Organizational Affiliation: