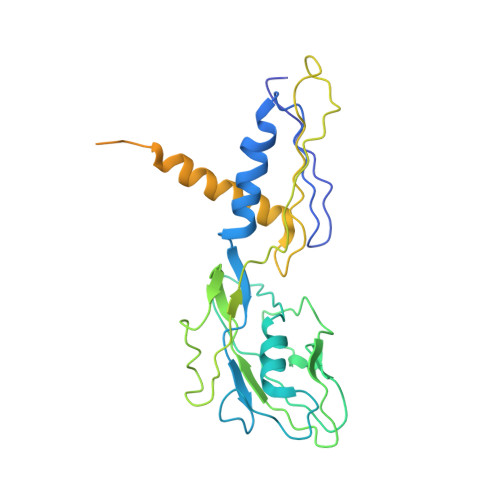
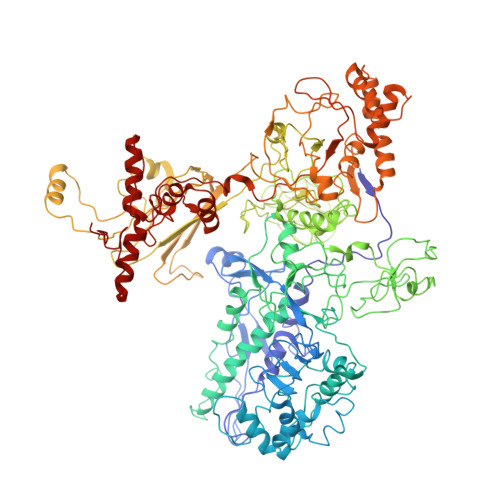
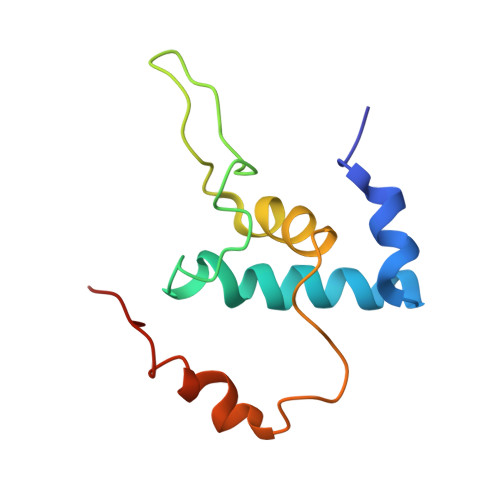
Bacterial RNA polymerase subunit omega and eukaryotic RNA polymerase subunit RPB6 are sequence, structural, and functional homologs and promote RNA polymerase assembly.
Minakhin, L., Bhagat, S., Brunning, A., Campbell, E.A., Darst, S.A., Ebright, R.H., Severinov, K.(2001) Proc Natl Acad Sci U S A 98: 892-897
- PubMed: 11158566 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.98.3.892
- Primary Citation Related Structures:
1HQM - PubMed Abstract:
Bacterial DNA-dependent RNA polymerase (RNAP) has subunit composition beta'betaalpha(I)alpha(II)omega. The role of omega has been unclear. We show that omega is homologous in sequence and structure to RPB6, an essential subunit shared in eukaryotic RNAP I, II, and III. In Escherichia coli, overproduction of omega suppresses the assembly defect caused by substitution of residue 1362 of the largest subunit of RNAP, beta'. In yeast, overproduction of RPB6 suppresses the assembly defect caused by the equivalent substitution in the largest subunit of RNAP II, RPB1. High-resolution structural analysis of the omega-beta' interface in bacterial RNAP, and comparison with the RPB6-RPB1 interface in yeast RNAP II, confirms the structural relationship and suggests a "latching" mechanism for the role of omega and RPB6 in promoting RNAP assembly.
- Waksman Institute, Department of Genetics, Department of Chemistry and Howard Hughes Medical Institute, Rutgers, The State University, Piscataway, NJ 08854, USA.
Organizational Affiliation: