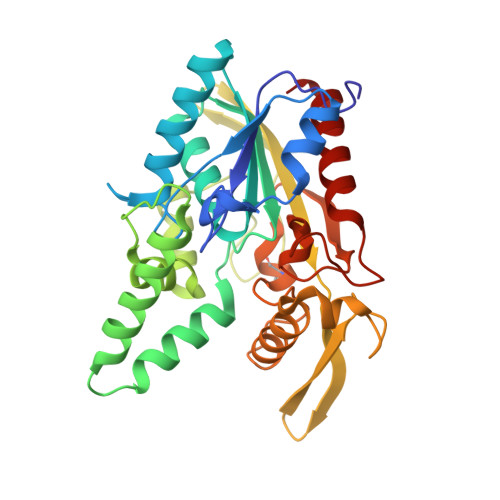
Complex of Burkholderia cepacia lipase with transition state analogue of 1-phenoxy-2-acetoxybutane: biocatalytic, structural and modelling study.
Luic, M., Tomic, S., Lescic, I., Ljubovic, E., Sepac, D., Sunjic, V., Vitale, L., Saenger, W., Kojic-Prodic, B.(2001) Eur J Biochem 268: 3964-3973
- PubMed: 11453990 Search on PubMed
- DOI: https://doi.org/10.1046/j.1432-1327.2001.02303.x
- Primary Citation Related Structures:
1HQD - PubMed Abstract:
In a series of four racemic phenoxyalkyl-alkyl carbinols, 1-phenoxy-2-hydroxybutane (1) is enantioselectively acetylated by Burkholderia cepacia (formerly Pseudomonas cepacia) lipase with an E value > or = 200, whereas for the other three racemates E was found to be < or = 4. To explain the high preference of B. cepacia lipase for (R)-(+)-1, a precursor of its transition state analogue with a tetrahedral P-atom, (R(P),S(P))-O-(2R)-(1-phenoxybut-2-yl)methylphosphonic acid chloride was prepared and crystallized in complex with B. cepacia lipase. The X-ray structure of the complex was determined, allowing to compare the conformation of the inhibitor with results of molecular modelling.
- Rudjer Boskovic Institute, Zagreb, Croatia] Institut für Chemie-Kristallographie, Freie Universität Berlin, Germany. luic@faust.irb.hr
Organizational Affiliation: