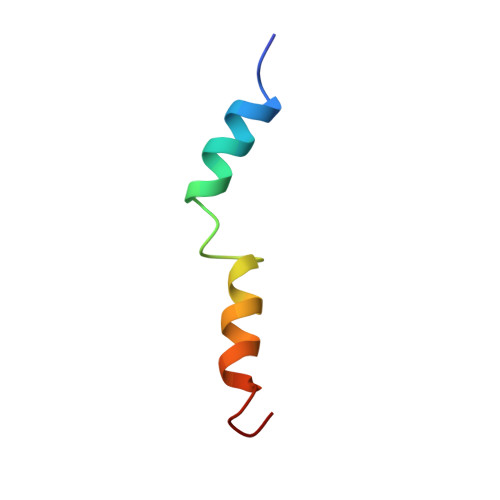
Solution structures of human parathyroid hormone fragments hPTH(1-34) and hPTH(1-39) and bovine parathyroid hormone fragment bPTH(1-37).
Marx, U.C., Adermann, K., Bayer, P., Forssmann, W.G., Rosch, P.(2000) Biochem Biophys Res Commun 267: 213-220
- PubMed: 10623601 Search on PubMed
- DOI: https://doi.org/10.1006/bbrc.1999.1958
- Primary Citation Related Structures:
1BWX, 1HPY, 1ZWA, 1ZWC - PubMed Abstract:
Parathyroid hormone (PTH) is involved in regulation of the calcium level in blood and has an influence on bone metabolism, thus playing a role in osteoporosis therapy. In this study, the structures of the human PTH fragments (1-34) and (1-39) as well as bovine PTH(1-37) in aqueous buffer solution under near physiological conditions were determined using two-dimensional nuclear magnetic resonance spectroscopy. The overall structure of the first 34 amino acids of these three peptides is virtually identical, exhibiting a short NH(2)-terminal and a longer COOH-terminal helix as well as a defined loop region from His14 to Ser17, stabilized by hydrophobic interactions. bPTH(1-37), which has a higher biological activity, shows a better-defined NH(2)-terminal part. In contrast to NH(2)-terminal truncations, which cause destabilization of helical structure, neither COOH-terminal truncation nor elongation significantly influences the secondary structure. Furthermore, we investigated the structure of hPTH(1-34) in 20% trifluoroethanol solution. In addition to its helix-stabilizing effect, trifluorethanol causes the loss of tertiary hydrophobic interactions.
- Lehrstuhl für Biopolymere, Universität Bayreuth, Bayreuth, D-95440, Germany. ute.marxf@uni-bayreuth.de
Organizational Affiliation: