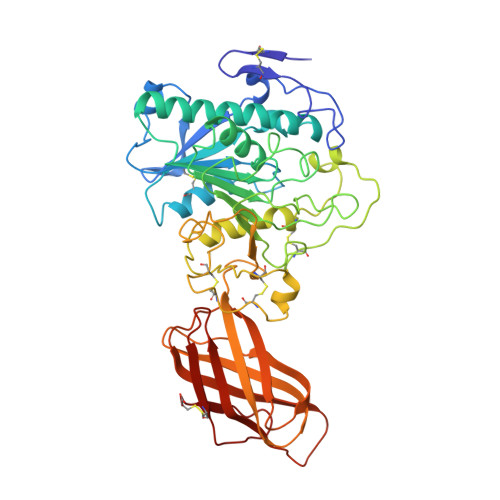
Horse pancreatic lipase. The crystal structure refined at 2.3 A resolution.
Bourne, Y., Martinez, C., Kerfelec, B., Lombardo, D., Chapus, C., Cambillau, C.(1994) J Mol Biology 238: 709-732
- PubMed: 8182745 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1331
- Primary Citation Related Structures:
1HPL - PubMed Abstract:
Pancreatic lipase (EC 3.1.1.3) plays a key role in dietary fat digestion by converting triacylglycerols into 2-monoacylglycerols and free fatty acids in the intestine. Although the crystallographic structures of the human pancreatic lipase and of a human lipase-porcine colipase complex have been solved, no refined structure of pancreatic lipase has yet been published. The crystal structure of the horse enzyme was solved by the molecular replacement method from the model of the human pancreatic lipase and subsequently refined to 2.3 A resolution. The final model contains two molecules of 449 amino acid residues each in the asymmetric unit, 705 well-defined water molecules and two calcium ions. The two molecules in the asymmetric unit of the orthorhombic crystals are related by a 2-fold non-crystallographic symmetry axis as in the case of the human lipase. However, the association between the two molecules in their respective crystal forms is different. The overall molecular structure of the horse lipase is very similar to that of the human enzyme. The horse lipase is made up of two well-defined domains. The N-terminal domain which bears the active centre has a typical alpha/beta hydrolase fold topology. The C-terminal domain which is devoted to colipase binding has a beta-sheet sandwich topology. Comparison of equivalent C alpha atom positions between the final model of the horse lipase and the human lipase structure shows only slight differences which are mainly located in the C-terminal domain. The horse enzyme possesses the common features of the known mammalian and microbial lipases, in particular the "flap" covering the catalytic triad. In addition to more precise information concerning these features, the elucidation of the horse lipase crystal structure allowed us to better understand the structural basis of the kinetic behaviour of pancreatic lipases towards a soluble substrate, p-nitrophenyl acetate, and the different sensitivity of these enzymes towards limited proteolysis.
- Laboratoire de Cristallographie et de Cristallisation des Macromolécules Biologiques CNRS, URA 1296, Faculté de Médecine Secteur-Nord, Marseille, France.
Organizational Affiliation: