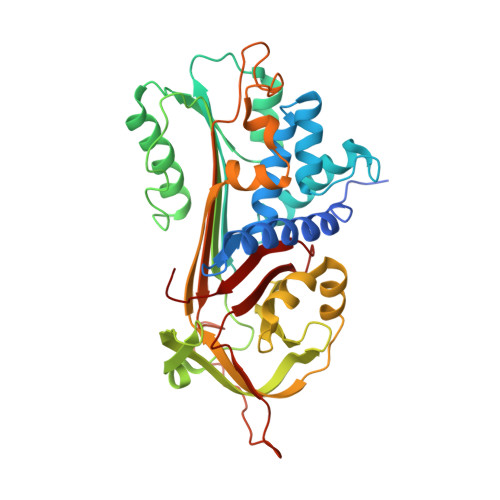
A 2.1 A resolution structure of an uncleaved alpha(1)-antitrypsin shows variability of the reactive center and other loops.
Kim, S., Woo, J., Seo, E.J., Yu, M., Ryu, S.(2001) J Mol Biology 306: 109-119
- PubMed: 11178897 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4357
- Primary Citation Related Structures:
1HP7 - PubMed Abstract:
Serpin (serine protease inhibitor) proteins are involved in diverse physiological processes including inflammation, coagulation, matrix remodeling, and cell differentiation. Deficiency of normal serpin functions leads to various hereditary diseases. Besides their clinical importance, serpin proteins draw much attention due to the large conformational changes that occur upon interaction with proteases. We present here the crystal structure of an uncleaved alpha(1)-antitrypsin determined by the multiple isomorphous replacement method and refined to 2.1 A resolution. The structure, which is the first active serpin structure based on experimental phases, reveals novel conformations in the flexible loops, including the proximal hinge region of the reactive center loop and the surface cavity region in the central beta-sheet, sheet A. The determined loop conformation explains the results of recent mutagenesis studies and provides detailed insights into the protease inhibition mechanism. The high-resolution structure of active alpha(1)-antitrypsin also provides evidence for the existence of localized van-der-Waals strain in the central hydrophobic core.
- Center for Cellular Switch Protein Structure, Korea Research Institute of Bioscience and Biotechnology, Yusong, Taejon, 305-600, Korea.
Organizational Affiliation: