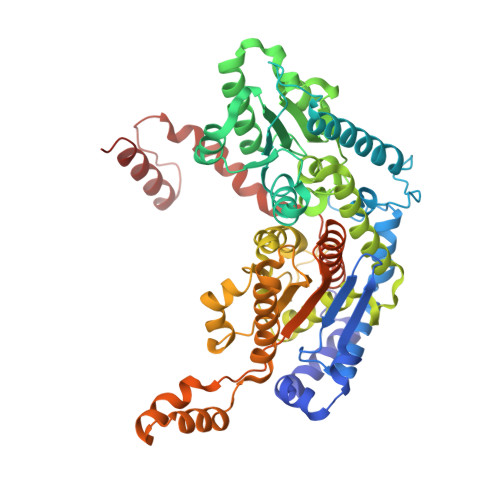
Crystal structure of rabbit phosphoglucose isomerase complexed with its substrate D-fructose 6-phosphate.
Lee, J.H., Chang, K.Z., Patel, V., Jeffery, C.J.(2001) Biochemistry 40: 7799-7805
- PubMed: 11425306 Search on PubMed
- DOI: https://doi.org/10.1021/bi002916o
- Primary Citation Related Structures:
1HOX - PubMed Abstract:
Phosphoglucose isomerase (PGI, EC 5.3.1.9) catalyzes the interconversion of D-glucose 6-phosphate (G6P) and D-fructose 6-phosphate (F6P) and plays important roles in glycolysis and gluconeogenesis. Biochemical characterization of the enzyme has led to a proposed multistep catalytic mechanism. First, the enzyme catalyzes ring opening to yield the open chain form of the substrate. Then isomerization proceeds via proton transfer between C2 and C1 of a cis-enediol(ate) intermediate to yield the open chain form of the product. Catalysis proceeds in both the G6P to F6P and F6P to G6P directions, so both G6P and F6P are substrates. X-ray crystal structure analysis of rabbit and bacterial PGI has previously identified the location of the enzyme active site, and a recent crystal structure of rabbit PGI identified Glu357 as a candidate functional group for transferring the proton. However, it was not clear which active site amino acid residues catalyze the ring opening step. In this paper, we report the X-ray crystal structure of rabbit PGI complexed with the cyclic form of its substrate, D-fructose 6-phosphate, at 2.1 A resolution. The location of the substrate relative to the side chains of His388 suggest that His388 promotes ring opening by protonating the ring oxygen. Glu216 helps to position His388, and a water molecule that is held in position by Lys518 and Thr214 accepts a proton from the hydroxyl group at C2. Comparison to a structure of rabbit PGI with 5PAA bound indicates that ring opening is followed by loss of the protonated water molecule and conformational changes in the substrate and the protein so that a helix containing amino acids 513-520 moves in toward the substrate to form additional hydrogen bonds with the substrate.
- Laboratory for Molecular Biology, MC567, Department of Biology, University of Illinois at Chicago, Chicago, Illinois 60607, USA.
Organizational Affiliation: