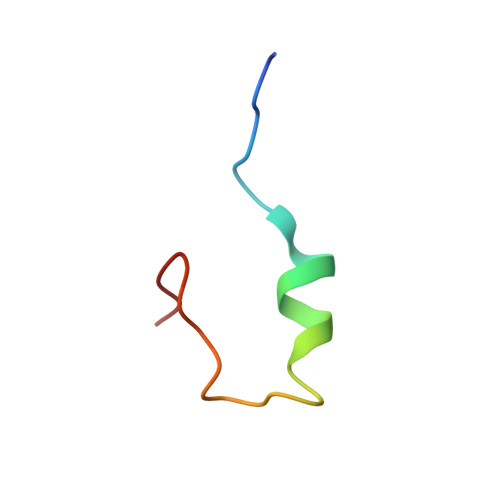
A new B-chain mutant of insulin: comparison with the insulin crystal structure and role of sulfonate groups in the B-chain structure
Dupradeau, F.Y., Richard, T., Le Flem, G., Oulyadi, H., Prigent, Y., Monti, J.P.(2002) J Pept Res 60: 56-64
- PubMed: 12081626
- DOI: https://doi.org/10.1034/j.1399-3011.2002.02990.x
- Primary Citation Related Structures:
1HO0 - PubMed Abstract:
The solution structure of a new B-chain mutant of bovine insulin, in which the cysteines B7 and B19 are replaced by two serines, has been determined by circular dichroism, 2D-NMR and molecular modeling. This structure is compared with that of the oxidized B-chain of bovine insulin [Hawkins et al. (1995) Int. J. Peptide Protein Res.46, 424-433]. Circular dichroism spectroscopy showed in particular that a higher percentage of helical secondary structure for the B-chain mutant is estimated in trifluoroethanol solution in comparison with the oxidized B-chain. 2D-NMR experiments confirmed, among multiple conformations, that the B-chain mutant presents defined secondary structures such as a alpha-helix between residues B9 and B19, and a beta-turn between amino acids B20 and B23 in aqueous trifluoroethanol. The 3D structures, which are consistent with NMR data and were obtained using a simulated annealing protocol, showed that the tertiary structure of the B-chain mutant is better resolved and is more in agreement with the insulin crystal structure than the oxidized B-chain structure described by Hawkins et al. An explanation could be the presence of two sulfonate groups in the oxidized insulin B-chain. Either by their charges and/or their size, such chemical groups could play a destructuring effect and thus could favor peptide flexibility and conformational averaging. Thus, this study provides new insights on the folding of isolated B-chains.
- GRBPD, UPRES 2629, Faculté de Pharmacie et de Médecine, Amiens, France.
Organizational Affiliation: