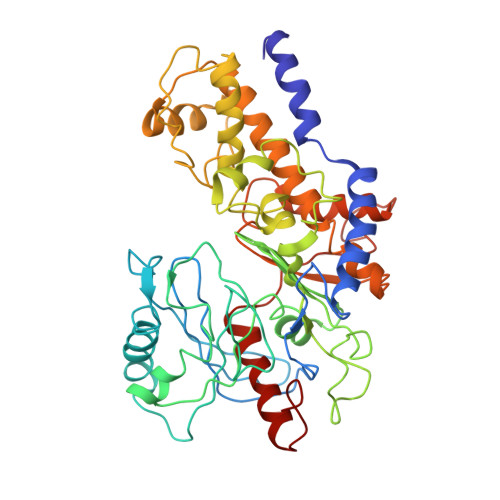
Structural dynamics of yeast hexokinase during catalysis.
Steitz, T.A., Shoham, M., Bennett Jr., W.S.(1981) Philos Trans R Soc London,ser B 293: 43-52
- PubMed: 6115422 Search on PubMed
- DOI: https://doi.org/10.1098/rstb.1981.0058
- Primary Citation Related Structures:
1HKG - PubMed Abstract:
The binding of the substrate glucose to yeast hexokinase results in a substantial enzyme conformational change that is essential for catalysis and may be important for the enzyme's specificity, as well as the control of its activity. From high-resolution crystal structures of the monomeric enzyme crystallized both in the presence and in the absence of glucose, we find that glucose binds into the deep cleft that separates the molecule into two lobes and causes these two lobes to move together and close off the cleft. The structure of the hexokinase crystallized in the presence of xylose and ADP is being determined at low resolution. In this crystal form, the enzyme was thought to be in the conformation of the ternary complex. However, a low-resolution structure of this crystal form shows clearly that the enzyme is in the 'open' form and is not a ternary complex. Crystals of the A isozyme with glucose and ADP may be. Further, chemically sequenced tryptic peptides are being incorporated into the model obtained by crystallographic refinement at 2.1 A resolution. Completion of the sequence and the structure of the ternary complex should allow a detailed description of the enzymatic mechanism of this kinase and the role of substrate-induced conformational changes in catalysis and control.