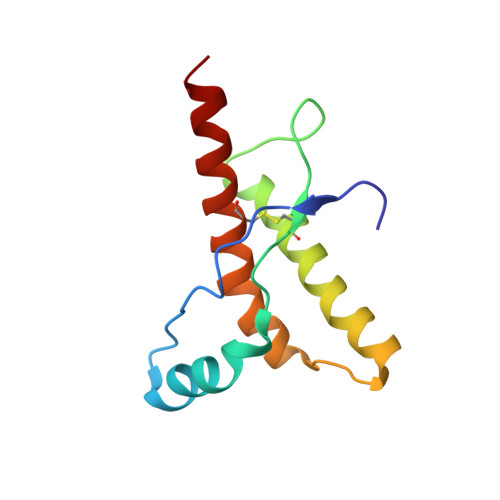
Influence of Ph on NMR Structure and Stability of the Human Prion Protein Globular Domain
Calzolai, L., Zahn, R.(2003) J Biological Chem 278: 35592
- PubMed: 12826672 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M303005200
- Primary Citation Related Structures:
1HJM, 1HJN - PubMed Abstract:
The NMR structure of the globular domain of the human prion protein (hPrP) with residues 121-230 at pH 7.0 shows the same global fold as the previously published structure determined at pH 4.5. It contains three alpha-helices, comprising residues 144-156, 174-194, and 200-228, and a short anti-parallel beta-sheet, comprising residues 128-131 and 161-164. There are slight, strictly localized, conformational changes at neutral pH when compared with acidic solution conditions: helix alpha1 is elongated at the C-terminal end with residues 153-156 forming a 310-helix, and the population of helical structure in the C-terminal two turns of helix alpha 2 is increased. The protonation of His155 and His187 presumably contributes to these structural changes. Thermal unfolding monitored by far UV CD indicates that hPrP-(121-230) is significantly more stable at neutral pH. Measurements of amide proton protection factors map local differences in protein stability within residues 154-157 at the C-terminal end of helix alpha 1 and residues 161-164 of beta-strand 2. These two segments appear to form a separate domain that at acidic pH has a larger tendency to unfold than the overall protein structure. This domain could provide a "starting point" for pH-induced unfolding and thus may be implicated in endosomic PrPC to PrPSc conformational transition resulting in transmissible spongiform encephalopathies.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Hönggerberg, CH-8093 Zürich, Switzerland. luigi@mol.biol.ethz.ch
Organizational Affiliation: