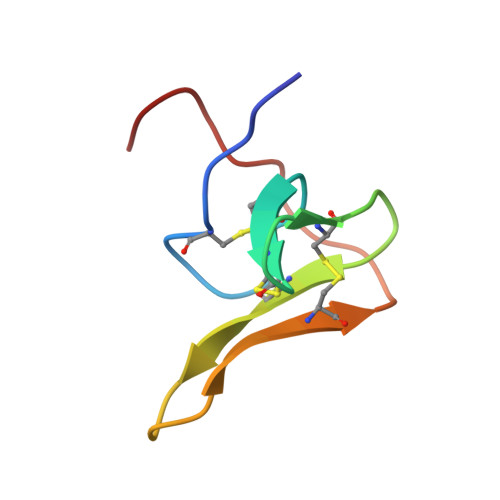
Nuclear magnetic resonance solution structure of hirudin(1-51) and comparison with corresponding three-dimensional structures determined using the complete 65-residue hirudin polypeptide chain.
Szyperski, T., Guntert, P., Stone, S.R., Wuthrich, K.(1992) J Mol Biology 228: 1193-1205
- PubMed: 1335515
- DOI: https://doi.org/10.1016/0022-2836(92)90325-e
- Primary Citation Related Structures:
1HIC - PubMed Abstract:
The three-dimensional structure of the N-terminal 51-residue domain of recombinant hirudin in aqueous solution was determined by 1H nuclear magnetic resonance (NMR) spectroscopy, and the resulting high-quality solution structure was compared with corresponding structures obtained from studies with the intact, 65-residue polypeptide chain of hirudin. On the basis of 580 distance constraints derived from nuclear Overhauser effects and 109 dihedral angle constraints, a group of 20 conformers representing the solution structure of hirudin(1-51) was computed with the program DIANA and energy-minimized with a modified version of the program AMBER. Residues 3 to 30 and 37 to 48 form a well-defined molecular core with two antiparallel beta-sheets composed of residues 14 to 16 and 20 to 22, and 27 to 31 and 36 to 40, and three reverse turns at residues 8 to 11 (type II), 17 to 20 (type II') and 23 to 26 (type II). The average root-mean-square deviation of the individual NMR conformers relative to their mean co-ordinates is 0.38 A for the backbone atoms and 0.77 A for all heavy atoms of these residues. Increased structural disorder was found for the N-terminal dipeptide segment, the loop at residues 31 to 36, and the C-terminal tripeptide segment. The solution structure of hirudin(1-51) has the same molecular architecture as the corresponding polypeptide segment in natural hirudin and recombinant desulfatohirudin. It is also closely similar to the crystal structure of the N-terminal 51-residue segment of hirudin in a hirudin-thrombin complex, with root-mean-square deviations of the crystal structure relative to the mean solution structure of 0.61 A for the backbone atoms and 0.91 A for all heavy atoms of residues 3 to 30 and 37 to 48. Further coincidence is found for the loop formed by residues 31 to 36, which shows increased structural disorder in all available solution structures of hirudin, and of which residues 32 to 35 are not observable in the electron density map of the thrombin complex. Significant local structural differences between hirudin(1-51) in solution and hirudin in the crystalline thrombin complex were identified mainly for the N-terminal tripeptide segment and residues 17 to 21. These are further analyzed in an accompanying paper.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule-Hönggerberg, Zürich, Switzerland.
Organizational Affiliation: