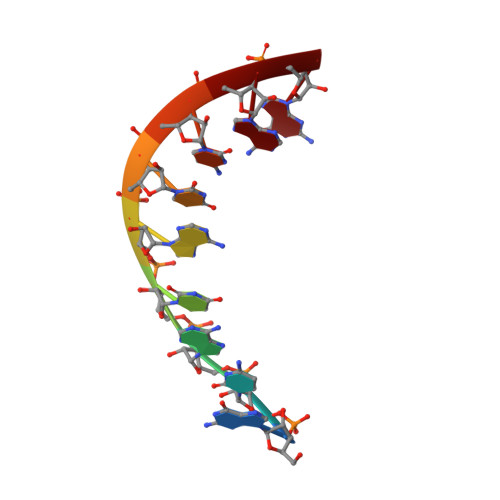
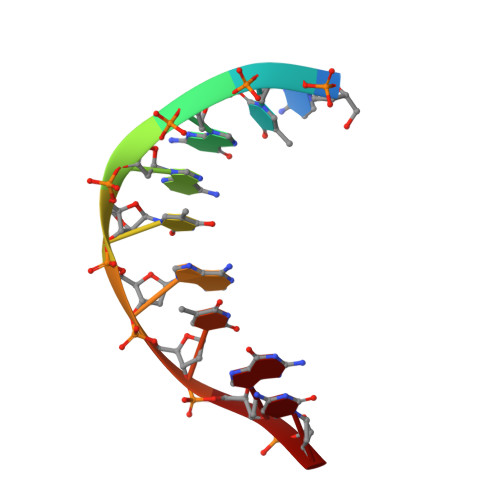
Locked Nucleic Acid (Lna) Recognition of RNA: NMR Solution Structures of Lna:RNA Hybrids
Petersen, M., Bondensgaard, K., Wengel, J., Jacobsen, J.P.(2002) J Am Chem Soc 124: 5974
- PubMed: 12022830 Search on PubMed
- DOI: https://doi.org/10.1021/ja012288d
- Primary Citation Related Structures:
1HHW, 1HHX - PubMed Abstract:
Locked nucleic acids (LNAs) containing one or more 2'-O,4'-C-methylene-linked bicyclic ribonucleoside monomers possess a number of the prerequisites of an effective antisense oligonucleotide, e.g. unprecedented helical thermostability when hybridized with cognate RNA and DNA. To acquire a detailed understanding of the structural features of LNA giving rise to its remarkable properties, we have conducted structural studies by use of NMR spectroscopy and now report high-resolution structures of two LNA:RNA hybrids, the LNA strands being d(5'-CTGAT(L)ATGC-3') and d(5'-CT(L)GAT(L)AT(L)GC-3'), respectively, T(L) denoting a modified LNA monomer with a thymine base, along with the unmodified DNA:RNA hybrid. In the structures, the LNA nucleotides are positioned as to partake in base stacking and Watson-Crick base pairing, and with the inclusion of LNA nucleotides, we observe a progressive change in duplex geometry toward an A-like duplex structure. As such, with the inclusion of three LNA nucleotides, the hybrid adopts an almost canonical A-type duplex geometry, and thus it appears that the number of modifications has reached a saturation level with respect to structural changes, and that further incorporations would furnish only minute changes in the duplex structure. We attempt to rationalize the conformational steering induced by the LNA nucleotides by suggesting that the change in electronic density at the brim of the minor groove, introduced by the LNA modification, is causing an alteration of the pseudorotational profile of the 3'-flanking nucleotide, thus shifting this sugar equilibrium toward N-type conformation.
- Nucleic Acid Center, Department of Chemistry, University of Southern Denmark, Odense University, DK-5230 Odense M, Denmark. mip@chem.sdu.dk
Organizational Affiliation: