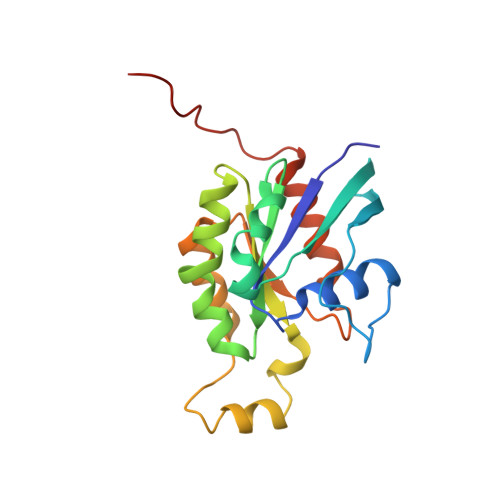
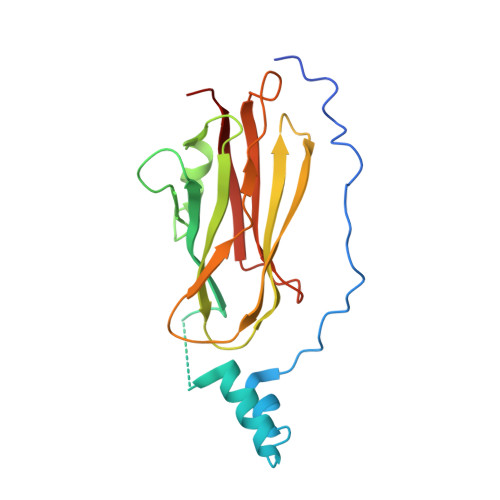
Crystal Structure of the Rac1-Rhogdi Complex Involved in Nadph Oxidase Activation
Grizot, S., Faure, J., Fieschi, F., Vignais, P.V., Dagher, M.-C., Pebay-Peyroula, E.(2001) Biochemistry 40: 10007
- PubMed: 11513578 Search on PubMed
- DOI: https://doi.org/10.1021/bi010288k
- Primary Citation Related Structures:
1HH4 - PubMed Abstract:
A heterodimer of prenylated Rac1 and Rho GDP dissociation inhibitor was purified and found to be competent in NADPH oxidase activation. Small angle neutron scattering experiments confirmed a 1:1 stoichiometry. The crystal structure of the Rac1-RhoGDI complex was determined at 2.7 A resolution. In this complex in which Rac1 is bound to GDP, the switch I region of Rac1 is in the GDP conformation whereas the switch II region resembles that of a GTP-bound GTPase. Two types of interaction between RhoGTPases and RhoGDI were investigated. The lipid-protein interaction between the geranylgeranyl moiety of Rac1 and RhoGDI resulted in numerous structural changes in the core of RhoGDI. The interactions between Rac1 and RhoGDI occur through hydrogen bonds which involve a number of residues of Rac1, namely, Tyr64(Rac), Arg66(Rac), His103(Rac), and His104(Rac), conserved within the Rho family and localized in the switch II region or in its close neighborhood. Moreover, in the switch II region of Rac1, hydrophobic interactions involving Leu67(Rac) and Leu70(Rac) contribute to the stability of the Rac1-RhoGDI complex. Inhibition of the GDP-GTP exchange in Rac1 upon binding to RhoGDI partly results from interaction of Thr35(Rac) with Asp45(GDI). In the Rac1-RhoGDI complex, the accessibility of the effector loops of Rac1 probably accounts for the ability of the Rac1-RhoGDI complex to activate the NADPH oxidase.
- Institut de Biologie Structurale, CEA-CNRS-UJF, UMR 5075, 41 Rue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: