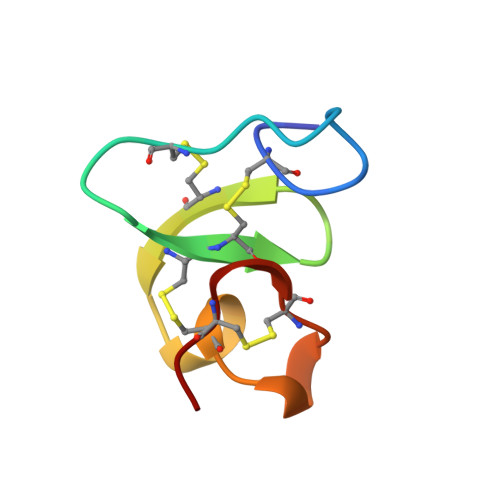
Hevein: NMR assignment and assessment of solution-state folding for the agglutinin-toxin motif.
Andersen, N.H., Cao, B., Rodriguez-Romero, A., Arreguin, B.(1993) Biochemistry 32: 1407-1422
- PubMed: 8431421 Search on PubMed
- DOI: https://doi.org/10.1021/bi00057a004
- Primary Citation Related Structures:
1HEV - PubMed Abstract:
The first high-resolution solution-state structure of a member of the toxin-agglutinin folding motif with the WGA disulfide linkage is presented. The 1H NMR spectrum of hevein has been 100% assigned from residue 2 through residue 43, the C-terminus, using two-dimensional correlation and NOE spectroscopy. During the course of the NOESY analysis, the three-dimensional structural features of hevein were derived, using nonstereospecific distance constraints (with tight bounds) for XPLOR simulated annealing followed by unconstrained relaxation in the CHARMm force field, at two levels of long-range constraint density. In addition, a large number of low-bound-only constraints, corresponding to unobserved NOE's, were used in both refinements. The first structure elucidation employed a total of 180 distance constraints (60 of which were medium or long range, i/i+n with n < or = 2). The second refinement employed 244 (101 medium or long range) constraints: some conformation-insensitive intraresidue constraints were deleted, two misassigned long-range constraints were corrected, and 41 new i/i+n (n > or = 2) constraints were added. The average bounds precisions of the two refinements were comparable (+/- 0.44 A) and significantly tighter than those that result when a universal low bound corresponding to the sum of the van der Waals radii was used. (The more conservative treatment of NOE's gave the same final structure but required a higher constraint density before assignment errors would stand out during the refinement.) Constraint density also has a significant influence on convergence and accuracy using tight constraints. The study demonstrates that convergence within an ensemble of solution structures is not a dependable criterion for either the accuracy or precision of the derived structure. The best fitting conformers from the refinement at the higher constraint density bear a greater similarity to the solid-state structure of the domains of wheat germ agglutinin (0.95 A rmsd over residues 2-32) than to the recently reported 2.8-A X-ray structure of hevein (1.25 A rmsd over residues 2-32, 2.83 A rmsd over residues 2-42). The consensus conformer from the solution data is defined to a backbone rmsd of < 0.6 A over the full sequence for which NMR data could be collected.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Chemistry, University of Washington, Seattle 98195.
Organizational Affiliation: