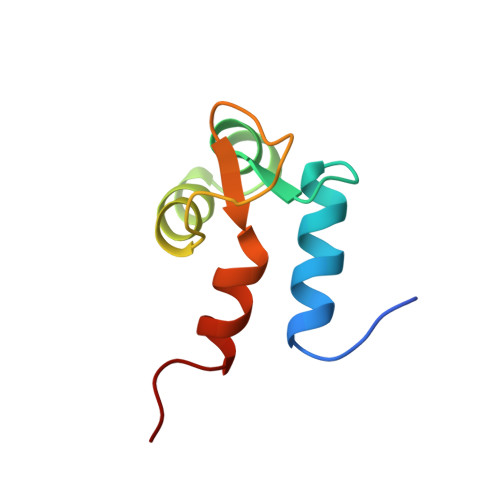
Ca2+-Independent Binding of an EF-Hand Domain to a Novel Motif in the Alpha-Actinin-Titin Complex
Atkinson, R.A., Joseph, C., Kelly, G., Muskett, F.W., Frenkiel, T.A., Nietlispach, D., Pastore, A.(2001) Nat Struct Biol 8: 853
- PubMed: 11573089 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1001-853
- Primary Citation Related Structures:
1H8B - PubMed Abstract:
The interaction between alpha-actinin and titin, two modular muscle proteins, is essential for sarcomere assembly. We have solved the solution structure of a complex between the calcium-insensitive C-terminal EF-hand domain of alpha-actinin-2 and the seventh Z-repeat of titin. The structure of the complex is in a semi-open conformation and closely resembles that of myosin light chains in their complexes with heavy chain IQ motifs. However, no IQ motif is present in the Z-repeat, suggesting that the semi-open conformation is a general structural solution for calcium-independent recognition of EF-hand domains.
- Division of Molecular Structure, National Institute for Medical Research, The Ridgeway, Mill Hill, London, NW7 1AA UK.
Organizational Affiliation: