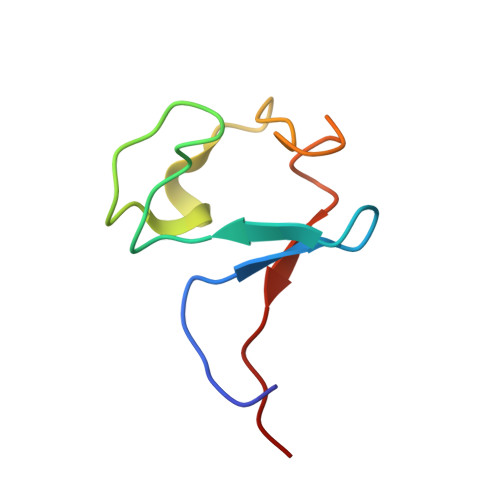
Solution structure of a zinc substituted eukaryotic rubredoxin from the cryptomonad alga Guillardia theta.
Schweimer, K., Hoffmann, S., Wastl, J., Maier, U.G., Rosch, P., Sticht, H.(2000) Protein Sci 9: 1474-1486
- PubMed: 10975569 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.9.8.1474
- Primary Citation Related Structures:
1DX8, 1H7V - PubMed Abstract:
The rubredoxin from the cryptomonad Guillardia theta is one of the first examples of a rubredoxin encoded in a eukaryotic organism. The structure of a soluble zinc-substituted 70-residue G. theta rubredoxin lacking the membrane anchor and the thylakoid targeting sequence was determined by multidimensional heteronuclear NMR, representing the first three-dimensional (3D) structure of a eukaryotic rubredoxin. For the structure calculation a strategy was applied in which information about hydrogen bonds was directly inferred from a long-range HNCO experiment, and the dynamics of the protein was deduced from heteronuclear nuclear Overhauser effect data and exchange rates of the amide protons. The structure is well defined, exhibiting average root-mean-square deviations of 0.21 A for the backbone heavy atoms and 0.67 A for all heavy atoms of residues 7-56, and an increased flexibility toward the termini. The structure of this core fold is almost identical to that of prokaryotic rubredoxins. There are, however, significant differences with respect to the charge distribution at the protein surface, suggesting that G. theta rubredoxin exerts a different physiological function compared to the structurally characterized prokaryotic rubredoxins. The amino-terminal residues containing the putative signal peptidase recognition/cleavage site show an increased flexibility compared to the core fold, but still adopt a defined 3D orientation, which is mainly stabilized by nonlocal interactions to residues of the carboxy-terminal region. This orientation might reflect the structural elements and charge pattern necessary for correct signal peptidase recognition of the G. theta rubredoxin precursor.
- Lehrstuhl für Biopolymere, Universität Bayreuth, Germany.
Organizational Affiliation: