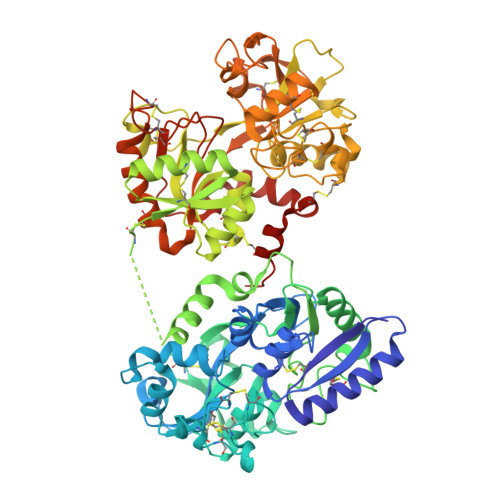
The Crystal and Molecular Structures of Diferric Porcine and Rabbit Serum Transferrins at Resolutions of 2.15 And 2.60A, Respectively
Hall, D.R., Hadden, J.M., Leonard, G.A., Bailey, S., Neu, M., Winn, M., Lindley, P.F.(2002) Acta Crystallogr D Biol Crystallogr 58: 70
- PubMed: 11752780 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444901017309
- Primary Citation Related Structures:
1H76, 1JNF - PubMed Abstract:
The serum transferrins are monomeric proteins with a molecular mass of around 80 kDa and are responsible for the transport of iron in vertebrates. The three-dimensional structures of diferric porcine and rabbit serum transferrin have been refined against X-ray diffraction data extending to 2.15 and 2.60 A, respectively. Data for both proteins were collected using synchrotron radiation at temperatures of 277 K. The porcine protein crystallizes in the space group C2, with unit-cell parameters a = 223.8, b = 44.9, c = 78.9 A, beta = 105.4 degrees with one molecule in the asymmetric unit. The structure was solved by molecular-replacement methods using rabbit serum transferrin as the search model. The structure was refined using REFMAC, with a final residual of 13.8% (R(free) = 18.2% for a 5% data sample) for all data to 2.15 A. The final model comprises 5254 protein atoms, two Fe(3+) cations and two CO(3)(2-) anions, one N-acetyl glucosamine moiety and 494 water molecules. The rabbit protein crystallizes in space group P4(3)2(1)2, with unit-cell parameters a = 127.2, c = 144.9 A and one molecule per asymmetric unit. The structure was solved using the method of multiple isomorphous replacement and refined using REFMAC to give a final residual of 18.6% (R(free) = 22.2% for a 5% data sample) for all data to 2.60 A. The final model comprises 5216 protein atoms, two Fe(3+) cations and two CO(3)(2-) anions, a Cl(-) anion and 206 solvent molecules; there is no clear indication of the carbohydrate moiety attached to Asn490 (rabbit serum numbering). Both molecules adopt a bilobal structure typical for members of the transferrin family. Each of the structurally homologous lobes contains two dissimilar domains with a single iron-binding site buried within the interdomain cleft. The porcine serum protein lacks an interdomain disulfide bridge close to the connecting peptide between the lobes, but this seems to have little effect on the overall orientation of the lobes. The N-lobes of both proteins possess lysine residues, one from each of the two domains, that lie in close proximity to one another to form the so-called dilysine trigger. The more acid-labile release of iron from serum transferrins than from lactoferrins is discussed.
- CLRC Daresbury Laboratory, Warrington, Cheshire WA4 4AD, England.
Organizational Affiliation: