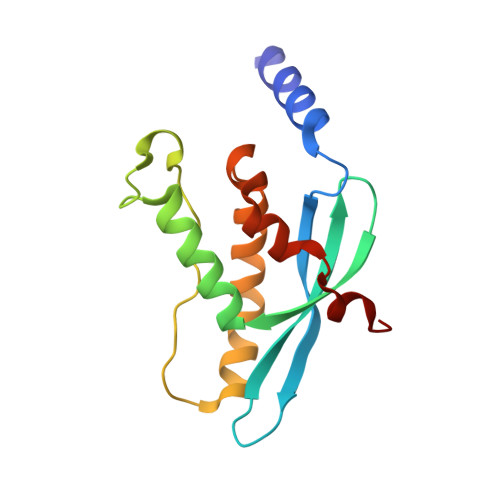
The Crystal Structure of the Px Domain from P40Phox Bound to Phosphatidylinositol 3-Phosphate
Bravo, J., Karathanassis, D., Pacold, C.M., Pacold, M.E., Ellson, C.D., Anderson, K.E., Butler, J.G., Lavenir, I., Perisic, O., Hawkins, P.T., Stephens, L., Williams, R.L.(2001) Mol Cell 8: 829
- PubMed: 11684018 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(01)00372-0
- Primary Citation Related Structures:
1H6H - PubMed Abstract:
More than 50 human proteins with a wide range of functions have a 120 residue phosphoinositide binding module known as the PX domain. The 1.7 A X-ray crystal structure of the PX domain from the p40(phox) subunit of NADPH oxidase bound to PtdIns(3)P shows that the PX domain embraces the 3-phosphate on one side of a water-filled, positively charged pocket and reveals how 3-phosphoinositide specificity is achieved. A chronic granulomatous disease (CGD)-associated mutation in the p47(phox) PX domain that abrogates PtdIns(3)P binding maps to a conserved Arg that does not directly interact with the phosphoinositide but instead appears to stabilize a critical lipid binding loop. The SH3 domain present in the full-length protein does not affect soluble PtdIns(3)P binding to the p40(phox) PX domain.
- Laboratory of Molecular Biology, Medical Research Council, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: