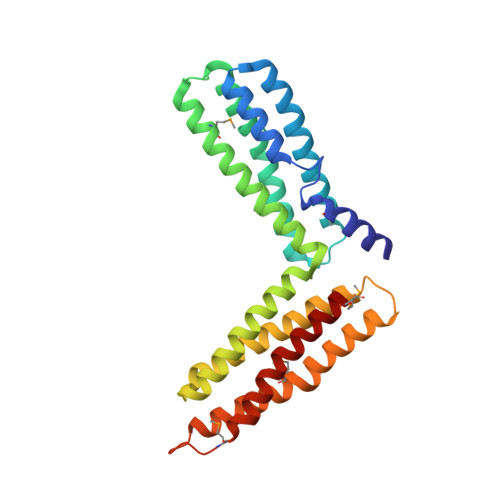
Crystal Structure of the M-Fragment of Alpha-Catenin: Implications for Modulation of Cell Adhesion.
Yang, J., Dokurno, P., Tonks, N.K., Barford, D.(2001) EMBO J 20: 3645
- PubMed: 11447106
- DOI: https://doi.org/10.1093/emboj/20.14.3645
- Primary Citation of Related Structures:
1H6G - PubMed Abstract:
The cytoskeletal protein alpha-catenin, which shares structural similarity with vinculin, is required for cadherin-mediated cell adhesion, and functions to modulate cell adhesive strength and to link the cadherins to the actin-based cytoskeleton. Here we describe the crystal structure of a region of alpha-catenin (residues 377-633) termed the M-fragment. The M-fragment is composed of a tandem repeat of two antiparallel four-helix bundles of virtually identical architectures that are related in structure to the dimerization domain of alpha-catenin and the tail region of vinculin. These results suggest that alpha-catenin is composed of repeating antiparallel helical domains. The region of alpha-catenin previously defined as an adhesion modulation domain corresponds to the C-terminal four-helix bundle of the M-fragment, and in the crystal lattice these domains exist as dimers. Evidence for dimerization of the M-fragment of alpha-catenin in solution was detected by chemical cross-linking experiments. The tendency of the adhesion modulation domain to form dimers may explain its biological activity of promoting cell-cell adhesiveness by inducing lateral dimerization of the associated cadherin molecule.
- Section of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.
Organizational Affiliation: