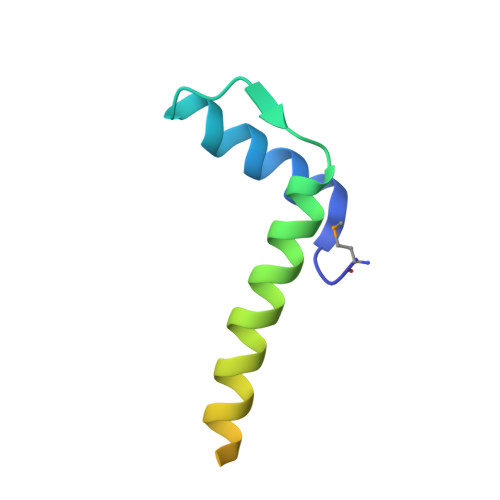
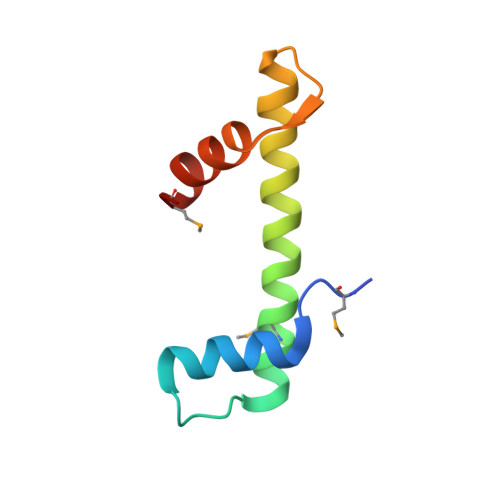
Crystal Structure of a Subcomplex of Human Transcription Factor TFIID Formed by TATA Binding Protein-Associated Factors Htaf4 (Htaf(II)135) and Htaf12 (Htaf(II)20).
Werten, S., Mitschler, A., Romier, C., Gangloff, Y.-G., Thuault, S., Davidson, I., Moras, D.(2002) J Biological Chem 277: 45502
- PubMed: 12237304 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M206587200
- Primary Citation Related Structures:
1H3O - PubMed Abstract:
The crystal structure is presented of a complex formed by the interacting domains from two subunits of the general transcription factor TFIID, the human TATA binding protein-associated factors hTAF4 (hTAF(II)135) and hTAF12 (hTAF(II)20). In agreement with predictions, hTAF12 forms a histone fold that is very similar to that of histone H2B, yet unexpected differences are observed between the structures of the hTAF12 interaction domain of hTAF4 and histone H2A. Most importantly, the hTAF4 fragment forms only the first two helices of a classical histone fold, which are followed by a 26-residue disordered region. This indicates that either full-length TAF4 contains an unusually long connecting loop between its second and third helix, and this helix is not required for stable interaction with TAF12, or that TAF4 represents a novel class of partial histone fold motifs. Structural models and structure-based sequence alignments support a role for TAF4b and hSTAF42/yADA1 as alternative partners for TAF12 and are consistent with the formation of nucleosome-like histone-fold octamers through interaction of TAF12 with a TAF6-TAF9 tetramer, yet argue against involvement of TAF12-containing histone-fold pairs in DNA binding.
- Institut de Génétique et de Biologie Moléculaire et Cellulaire, CNRS/INSERM/Université Louis Pasteur, Boîte Postale, B.P. 163 67404 Illkirch Cédex Communauté Urbaine de Strasbourg, France.
Organizational Affiliation: